-
Paper Information
- Previous Paper
- Paper Submission
-
Journal Information
- About This Journal
- Editorial Board
- Current Issue
- Archive
- Author Guidelines
- Contact Us
Clinical Medicine and Diagnostics
p-ISSN: 2163-1433 e-ISSN: 2163-1441
2014; 4(1A): 15-20
doi:10.5923/s.cmd.201401.03
Paraoxonase Activity and Gene Polymorphism in Rheumatoid Arthritis among Egyptians
Fathy. M. Elfasakhany1, 2, Saad A. Abou-Elnoeman1, Mervat E. Hussein3, Hanadi A. Lamfoon2, Mohamed M. Beyari2
1Department of Medical Biochemistry, Faculty of Medicine, Tanta University, Tanta, Egypt
2Department of Basic and Clinical Oral Sciences, Faculty of Dentistry, Umm Al Qura University, Saudi Arabia
3Department Physical Medicine and Rehabilitation, Faculty of Medicine, Tanta University, Tanta, Egypt
Correspondence to: Fathy. M. Elfasakhany, Department of Medical Biochemistry, Faculty of Medicine, Tanta University, Tanta, Egypt.
| Email: |  |
Copyright © 2012 Scientific & Academic Publishing. All Rights Reserved.
Paraoxonase1 (PON1) is an esterase enzyme that has antioxidant activity and is tightly associated with high density lipoprotein (HDL) in blood. Decreased activity of PON1 has been reported in many diseases and could be partially attributed to PON1 glutamine 192 arginine polymorphism. The aim of this study was to determine the paraoxonase (PON1) status, i.e.PON1 activities and phenotypes (QQ, QR and RR), and its relationship with lipid status and lipid peroxidation among Egyptians with rheumatoid arthritis (RA). The study included 40 RA patients and 40 healthy volunteers. The level of malondialdehyde (MDA), as a marker for oxidative stress, plasma paraoxonase 1 (PON1) and arylesterase (ARE) activities were determined using an enzymatic spectrophotometric method. PON1 192 gene polymorphism was determined using polymerase chain reaction based restriction fragment analysis. The results of the present work showed that PON1 and ARE activities were statistically lower (P <0.001) while the MDA level was found to be statistically higher (P<0.001) in the RA group compared with the control group. There was no significant difference in the genotype distribution between RA group and control group. No significant difference was observed between the levels of total triglycerides, total cholesterol, HDL-C and LDL-C in both groups. We concluded that PON1 and ARE activities, which have antiatherogenic capability, are decreased in patients with RA. PON1 and ARE activities may be affected by oxidative stress demonstrated by high MDA level which contribute to the pathogenesis of RA. PON1 192 Q/R polymorphism may not be associated with RA incidence in Egyptians.
Keywords: Rheumatoid arthritis, Oxidative stress, Paraoxonase, Polymorphism, MDA, Arylesterase, Lipid status
Cite this paper: Fathy. M. Elfasakhany, Saad A. Abou-Elnoeman, Mervat E. Hussein, Hanadi A. Lamfoon, Mohamed M. Beyari, Paraoxonase Activity and Gene Polymorphism in Rheumatoid Arthritis among Egyptians, Clinical Medicine and Diagnostics, Vol. 4 No. 1A, 2014, pp. 15-20. doi: 10.5923/s.cmd.201401.03.
Article Outline
1. Introduction
- Rheumatoid arthritis is a systemic disorder of unknown cause affecting 1 - 2% of the adult population with variation across regions of the world. Women are two to three times more likely to get RA than men, and for both sexes risk increases with age[1].The causes of RA have not been completely elucidated. Reactive oxygen species (ROS), such as superoxide radical, hydrogen peroxide, hydroxyl radical and hypochlorous acid contribute significantly to tissue injury in RA[2]. In cases of excess ROS or decreased level of antioxidant activity, including antioxidant enzymes especially paraoxonase, oxidative stress causes severe metabolic malfunctions, damage of cell membrane and further decomposition of peroxidised lipids results in production of a variety of end products including malondialdehyde (MDA)[3-5].The paraoxonase (PON) gene family consists of three members- PON1, PON2, and PON3-located adjacent to each other on the long arm of chromosome 7 in humans. These three human PON genes share approximately 60% identity at the amino acid level and approximately 70% identity at the nucleotide level[6].Human serum paraoxonase (PON 1:aryldialkylphosphatase[EC 3.1.8.1] is a calcium dependent glycoprotein of 354 amino acids with a molecular mass 43 kDa that is tightly associated with apolipoprotein A-1 in HDL, hydrolyzes carboxylic acid esters, organophosphate insecticides and nerve gases[7,8].PON1 is synthesized in the liver and secreted in the blood where it associates with high-density lipoprotein[9]. The enzyme decreases accumulation of the lipid peroxides in low density lipoprotein (LDL) due to its ability to reduce hydroperoxides and attenuates biological effects of mildly oxidized LDL[10].Serum paraoxonase 1 activity appears to be different among individuals and populations. The molecular basis of such variations may be due to polymorphism in paraoxonase 1 gene[12]. Paraoxonase 1 has two polymorphisms at the coding region, one in position 192, which is a glutamine/arginine substitution and the other at position 55 which is a methionine/leucine substitution[12,13]. Thesepolymorphisms may affect the hydrolytic activity of PON 1 isoenzymes with respect to lipid peroxidation[14,15].It has been reported that paraoxonase activity of the glutamine 192 (Q allele) and methione 55 (M allele) isoforms is lower than that of the arginine 192 (R allele) and leucine 55 (L allele) isoforms[14-16]. Nevertheless, PON1 in QQ and MM genotype subjects seems to be more effective in protecting LDLs from oxidation than that in RR and LL genotype subjects[15].The relationship between PON1 activity and PON1 192 polymorphism in rheumatoid arthritis is still not well investigated. So, the aim of the present study was to shed a light on the role of PON1 activity and PON1 192 polymorphism with rheumatoid arthritis among Egyptians.
2. Material and Methods
2.1. Subjects
- Patients in the present study were all outpatients attending the Department of Physical Medicine & Rehabilitation, Tanta University Hospital, Tanta, Egypt. All patients were Rheumatoid arthritis patients diagnosed according to American Rheumatism Association (ARA) revised criteria. They were classified according to the activity of the disease into 32 inactive and 8 active. Patients with D.M., renal, hepatic diseases or coronary artery disease were excluded from the study. Fully informed consent was obtained. The control subjects consisted of 40 healthy subjects who either attended a routine health check at a general practice or at their place at work. Venous blood was collected from all subjects between 9:00 and 11:00 a.m. after fasting from 10:00–11:00 p.m. the previous day. Each sample was divided into two halves, one half for serum preparation and the other half was put in sterile K3EDTA (tri-potassium ethylenediaminetetraacetic acid) coated tubes were used for the DNA extraction samples. Plasma was isolated by low speed centrifugation, white cells were removed from the buffy coat for DNA extraction. Samples were stored at –20℃ till the time of use.
2.2. Determination of Serum PON1 Activity
- PON1 activity was measured by adding plasma to Tris buffer (100 mmol/l, pH 8.0) containing 2 mmol/l CaCl2 and 1 mmol/l paraoxon (O, O-diethyl-O-nitrophenylphosphate (Sigma) The rate of generation of P-nitrophenol was determined at 405 nm, 25℃ with the use of continuously recording spectrophotometer as described previously[16,17].
2.3. Determination of Serum Arylesterase Activity
- Arylesterase activity was measured using phenylacetate as a substrate as previously described[18]. The reaction mixture contained 750 μl of 0.1 mol/l Tris-HCl (pH 8.5), 1 mmol/l CaCl2, 125 µl of 12 mmol/l phenylacetate and 125 µl of diluted serum (1:10 diluted with water). Initial rates of hydrolysis were determined by following the increase of phenol concentration at 270 nm at 37°C on a CE 7250 spectrophotometer (Cecil Instruments Limited, UK). Enzyme activities were expressed in international units per 1 liter of serum (U/l). An international unit is the amount of hydrolyzed substrate in mmol/minute.
2.4. Measurement of Malondialdehyde (MDA)
- MDA level was measured according to a method described elsewhere[19]. That method depends on the fact that MDA reacts with thiobarbituric acid (TBA) producing thiobarbituric acid reactive substance (TBARS), a pink chromogen, which can be measured spectrophotometrically at 532 nm.MDA was expressed as nmol/ml and was calculated using 1.65 × 105 M-1 cm-1as molar absorption coefficient.
2.5. Determination of Plasma Lipids
- Plasma total cholesterol, HDL-C, were determined using CHOD-POD method (Spinreact, Santa Colma, Spain). Plasma triglycerides were measured by the enzymatic GPO-PAP method (Human, Wiesbaden, Germany). LDL-C was estimated using Friedwald formula: LDL-C = total cholesterol – (HDL cholesterol + triglycerides/5) mg/dl.
2.6. Detection of Rheumatoid Factor (RF) and C-reactive Protein (CRP)
- Rheumatoid factor was determined using rapid latex particles method (Spinreact, Santa Colma, Spain). C-reactive protein (CRP) was measured using semiqualitative method which is rapid and sensitive (Spinreact, Santa Colma, Spain).
2.7. Determination of PON1 Genotypes
- DNA was extracted from peripheral leukocytes using sodium dodecyl sulphate (SDS) lysis and ethanol precipitation[20]. Aliquots of genomic DNA were used for PCR amplification as described previously[13,21]. Two primers, the sense primer: 5` TAT TGT TGC TGT GGG ACC TGA G 3` and antisense primer: 5` CAC GCT AAA CCC AAA TAC ATC TC 3` were used for PCR amplification of 99 bp DNA fragment covering the region containing Gln192 or Arg 192. The PCR reaction mixture contained about 200 ng DNA template, 0.5 μM of each primer, 1.5 mM MgCl2, 200 μM of the 4 dNTP and 1 U Taq DNA polymerase (Amersham, Bioscience). After denaturing the DNA for 4 minutes at 95℃, the reaction mixture was subjected to 35 cycles of denaturing for 1 minute at 95℃, 1 minute annealing at 60℃ and 1 minute extension at 72℃. The 99 bp PCR product was digested with 8 U Alw1 restriction endonuclease (MBI Fermentas) overnight at 37℃ and the digested products were separated by electrophoresis on 2% agarose electrophoresis and visualized using ethidium bromide. The Arg genotype contains a unique Alw1 restriction site which results in 66 and 33 bp product, whereas the Gln genotype is not cleaved by this restriction enzyme.
2.8. Statistical Analyses
- data were analyzed using SPSS for Windows version 10.0 (SPSS Inc, Chicago. IL, USA). Student's t test, ANOVA and Scheffe post hoc test were used to compare mean values of continuous variable in cases and control, whereas χ2 analysis was used to compare categorical data. Relative risk was performed to examine genotype risk contribution.
3. Results
3.1. Serum Paraoxonase Activity
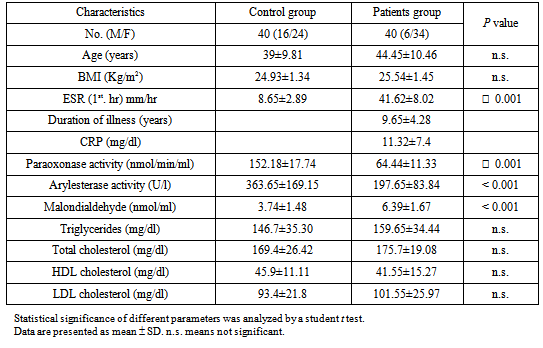
- Serum Paraoxonase activity was significantly decreased in rheumatoid arthritis subjects if compared with the control group (Table 1).
|
|
3.2. MDA Level
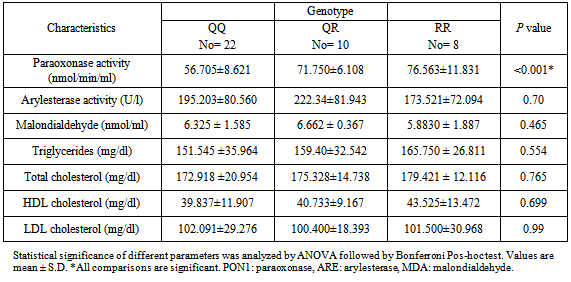
- There was a highly significant MDA level in rheumatoid arthritis group than control group as shown in Table 1. When correlated to genotypes, there was no significant difference in MDA level among the different genotypes in the patient group. (Table 4).
|
|
4. Discussion
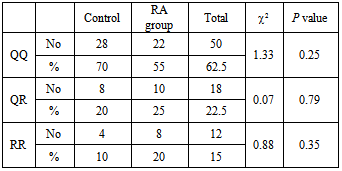
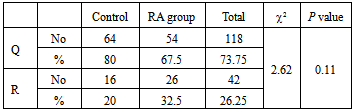
- Rheumatoid arthritis (RA) is the most common single cause of chronic synovitis, affecting multiple diarthroidial joints in a characteristic distribution, and leading to pain, deformities and a reduced quality of life. The precise aetiology of RA has not been established, but epidemiologic data as well as the current genetic data clearly reflect a complex genetic component, along with a largely unknown, and presumably variable contributions by environmental factors[22].Rheumatoid factor, which is an IgM directed against other immunoglobulins, are found in more than two-thirds of adult patients with RA, but they are not specific to RA and are found in patients with a number of other conditions. Additionally, five percent of otherwise healthy people have circulating rheumatoid factors. While the presence of rheumatoid factor is not useful as a screening test, high levels in combination with a suggestive clinical presentation of RA are prognostic for more severe disease with extra-articular manifestations[23]. All RA patients in the present study were RF positive. The disagreement between our results and results of other authors may be due to small number of the cases used in this study.Serum C-reactive protein concentration has been widely adopted as a marker of systemic inflammation. In RA, the level of CRP is frequently used in conjunction with assessments of articular swelling and tenderness to estimate the level of disease activity and, in addition to the erythrocyte sedimentation rate, is a component of the American College of Rheumatology core set for measuring clinical response, the Disease Activity Score and as a biomarker of structural damage in RA[24]. The present study reported high levels of CRP in our entire patient group. Eight patients in our study have shown the maximal level of CRP and ESR with severe clinical manifestations; they were considered to be active while the remaining patients were considered inactive. Several groups have demonstrated increased oxidative enzyme activity along with decreased antioxidant levels in RA sera and synovial fluids[25-28]. Because of the highly reactive nature of ROS, it is difficult to directly demonstrate their presence in vivo. It is considerably more practical to measure the ‘footprints’ of ROS, such as their effects on various lipids, proteins, and nucleic acids[29]. Lipid peroxidation is a ROS-mediated process that leads to cell membrane damage, resulting in cellular dysfunction or death. Lipid peroxides, derived from polyunsaturated fatty acids, are unstable and decompose to form a complex series of compounds. Identification and quantification of one or more of these end products can be utilized for the assessment of ROS-production in the inflamed joint. Several of these end product aldehydes, especially malondialdehyde (MDA), react with thiobarbituric acid (TBA) under acidic conditions to produce a pink-colored chromogen that absorbs light strongly at a wavelength of 532 nm, enabling a simple spectrophotometric measurement of lipid peroxidation 30. In the present study, MDA level was significantly elevated in RA patients. Similar results have been reported by other investigators[5,31-37].In the present work, lipid profile was determined because there is a close relationship between paraoxonase 1 (PON-1) and HDL. PON1 has recently emerged as the component of HDL most likely to explain its ability to metabolize lipid peroxides and to protect against their accumulation on LDL. The association of PON-1 with high-density lipoproteins (HDL) is a prerequisite for maintaining normal serum activity of the enzyme[38]. Total plasma cholesterol, triglycerides, HDL-C and LDL-C levels showed no significant difference between the control and the RA subjects. The results of this study were in basic agreement with Tanimotoet al[39],while Votteryet al[40]demonstrated that there is an inverse relationship between RA activity and lipid levels. The discrepancies in the values of lipid profile observed in the various studies might be due to differences in studied populations as well as in the disease activity.Furthermore, in the present study, PON-1 and Arylesterase (ARE) activities were decreased significantly in RA group compared with the control group. This corresponds to the previous reports of Baskolet al[35], Tanimotoet al[39], Altinadgeet al[41] and Isiket al[42]. The decrease in serum PON1 and ARE activity in RA patients might result from the increased inactivation of PON-1 and ARE due to the increased generation of reactive oxygen species in RA[43]. The second possible explanation may be due to a reduction of PON-1 and ARE independently of HDL-C concentration in RA which was shown by the result of the present study. The third possible explanation is that the decreased PON -1 activity may result from a reduction of PON1 production by the liver[39]. The PON1 Gln192Arggene polymorphism may alter significantly an individual’s susceptibility towards different substrates. It was found that Q (Gln) carriers have significantly lower serum PON1 activity than R (Arg) carriers. In the present study the QQ, QR and RR genotypes distribution were found to be 0.7, 0.1 and 0.2 respectively among the control group.This is slightly higher if compared to Caucasian populations, but significantly lowers than other Oriental populations. The difference in genotype distribution of PON-1 among Egyptian and other races may be due to different ethnic groups. Furthermore, we found that there was no significant difference in the distribution of Q and R alleles between the RA patients and healthy controls. Therefore, PON1 Q192R polymorphism might not be associated with the incidence of rheumatoid arthritis among Egyptian population. Tanimotoet al[39] found that there is a difference in the distribution of PON1 192 Q/R genotypes among RA Japanese patients and healthy subjects and serum PON1 activity with PON1-Q in RA patients was obviously low compared with that in healthy subjects despite the absence of differences in PON1-R between the two groups. This discrepancy might be due to the difference in ethnic populations. In the present study the QQ genotypes were the major prevailing type in the patient group with a 55% percentage followed by RR (25%) then QR (20%). The Q allele frequency was significantly higher than the R allele in both RA and control groups. The Q allele carriers were previously reported to be protected against peroxidation more than R allele carriers. In conclusion, serum PON1 activity decreased independently of PON1 192 polymorphism.The present study showed significant difference in PON-1 activity towards paraoxon in different genotypes. RR genotypes were the most active towards paraoxon and QQ genotypes were the least active while the activity of QR genotypes were intermediate. Moreover, there was no difference in arylesterase enzyme activity towards p-nitrophenyl acetate among different genotypes in RA patients. Tripiet al[42] studied the effect of PON1 192 polymorphism in systemic lupus erythematosus. They reported that The PON1 Q192R polymorphism has substantial influence on PON1 activity towards paraoxon, which has been reported in almost all published studies. In that study, despite the significant association between PON Q192R polymorphism and PON1 activity, there was no clear-cut association of PON1 Q192R polymorphism with SLE risk. Sidotiet al[43] reported that there is a correlation between PON1 Q192R polymorphism and multiple sclerosis (MS). On the other hand, HashemiM et al[46] had found no significant differences between rheumatoid arthritis patients and control subjects regarding PON1 Q192R polymorphism. PON1 Q192R polymorphism was not found to be correlated with increased risk for rheumatoid arthritis in this Iranian population. On the other hand, Charles-Schoeman C. et al[47] suggested a relationship between the genetic determinants and activity of PON1 with cardiovascular risk in RA patients as assessed by the presence or absence of carotid plaque. This discrepancy might be due to the difference in ethnic group.
5. Conclusions
- We concluded that PON1 activity is more important than PON1 polymorphism in rheumatoid arthritis. Our future plan is to study the PON1 polymorphisms in a larger population which may clarify the exact relationship between PON-1 polymorphism and disease activity in RA among Egyptian population.
ACKNOWLEDGEMENTS
- We thank Dr. M. Mona for his helpful comments and discussion and Zeinab Riad for her secretarial work. Fathy M. Elfasakhany is on secondment, Tanta University Faculty of Medicine, Medical Biochemistry Department, Tanta, Egypt.
 Abstract
Abstract Reference
Reference Full-Text PDF
Full-Text PDF Full-text HTML
Full-text HTML