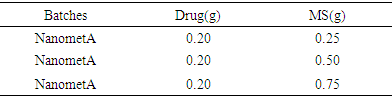Tenderwealth Clement Jackson1, Bernard Opatimidi Patani2, Nonye Linda Ifekpolugo1, Edidiong Michael Udofa1, Nkem Mercy Obiakor1
1Department of Pharmaceutics and Pharmaceutical Technology, University of Uyo, Uyo, Nigeria
2Medical Department, Agip Oil Company, Port-Harcourt, Rivers State, Nigeria
Correspondence to: Tenderwealth Clement Jackson, Department of Pharmaceutics and Pharmaceutical Technology, University of Uyo, Uyo, Nigeria.
| Email: |  |
Copyright © 2019 The Author(s). Published by Scientific & Academic Publishing.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/

Abstract
Development of Metronidazole loaded silver nanoparticles from Acalypha ciliata was carried out using the juice from the leaves of Acalypha ciliata as reducing agent and modified starch as stabilizing agent. Nanoparticles synthesized were labelled as nanometA, nanometB and nanometC. The percentage yield of the nanoparticles were above 88%. The entrapment efficiencies were 78.60%, 84.92% and 87.19% while the loading capacities were 26.18%, 12.10% and 7.92% for nanometA, nanometB and nanometC respectively. Differential scanning calorimetry (DSC) for optimized batch indicated the absence of interaction between the starch and drug. The UV VIS spectroscopy showed Surface Plasmon resonance of 407 nm for optimized batch while the mean particle size was 425.7 nm. All the batches showed controlled release of the drug from the Nanoparticles. The kinetics of release was mainly zero order for all the nanoparticles. Antimicrobial property of the optimized nanoparticle was significantly (P< 0.01) higher with increasing polymer to drug ratio.
Keywords:
Green synthesis, Nanoparticles, Starch, Characterisation, Antimicrobial and metronidazole
Cite this paper: Tenderwealth Clement Jackson, Bernard Opatimidi Patani, Nonye Linda Ifekpolugo, Edidiong Michael Udofa, Nkem Mercy Obiakor, Developent of Metronidazole Loaded Silver Nanoparticles from Acalypha ciliata for Treatment of Susceptible Pathogens, Nanoscience and Nanotechnology, Vol. 9 No. 1, 2019, pp. 22-28. doi: 10.5923/j.nn.20190901.02.
1. Introduction
Bio-synthesis of nanoparticles is a fast growing research in the field of nanotechnology. Extensive research has been carried out on silver nanoparticles as an important group of nanomaterials due to their peculiar biological, optical and physio-chemical properties [1].Nanoparticles can be synthesized easily by using various physical and chemical methods. Chemical reduction of metal salts using various reducing agents in the presence of stabilizer is currently of interest in the preparation of silver nanoparticles. Reducing agents such as sodium Borohydride (NaBH4), hydrazine (N2H4), formaldehyde, etc. can be used to reduce a silver containing salt to produce silver nanoparticles [2].But most of the chemical methods used for the synthesis of nanoparticles involve the use of toxic hazardous chemicals that create biological risk and sometimes these chemical processes are not eco-friendly. Therefore, there is a growing need to develop cost effective, non-toxic and eco-friendly methods for the synthesis of silver nanoparticles using simple techniques and readily available equipment. The use of plants and microorganisms in the synthesis of nanoparticles emerge as an eco-friendly and exciting approach [2,3]. In recent times, plant extract have been used as reducing and capping agent for the synthesis of nanoparticles. The use of plant extract is more beneficial as it does not involve sophisticated processes such as intracellular synthesis and multiple purification steps or the maintenance of microbial cell culture [4]. Nanoparticles are now considered viable alternatives to antibiotics as they seem to possess a high potential to address the problem of the emergence of bacterial multidrug resistance [5]. Silver nanoparticles have attracted much attention in the science [6]. Silver has always been used against various infections and functions as both antiseptic and antimicrobial agent against Gram-positive and Gram-negative bacteria [7]. Silver nanoparticles were considered, in recent years, particularly attractive for the production of a new class of antimicrobials [8,5,9], opening up a completely new way to combat a wide range of pathogenic bacteria.Silver nanoparticles have been synthesized with success using leaf extract from Carica papaya, Capsicum annum to mention a few. The choice of the plants used to synthesize nanoparticles is often based on the added biological value of the plant material. Acalypha ciliata has been reported to possess significant antimicrobial (antibacterial and antifungal) properties hence its use in this study.
2. Materials and Method
MATERIALS:AgNO3 (Sigma Aldrich, USA) and Acalypha ciliata were the materials used. Other chemicals and reagents were of laboratory grade.METHOD:A. PREPARATION OF METRONIDAZOLE LOADED SILVER NANOPARTICLES- Plant materials collection and processingFresh leaves of Acalypha ciliata were collected from a local farm in Uyo, Akwa Ibom State. The plant was identified and authenticated in the Department of Pharmacognosy and Natural Medicine, University of Uyo. The plant leaves were thoroughly washed with tap water to rinse off dusts and other unwanted materials accumulated on the leaves from their natural environment. The dust free leaves were allowed to dry under shade in the Pharmaceutics laboratory for 24 h. The dried leaves will be powdered by using electric blender.- Extraction procedure50 g of the powdered plant material were kept in 500 mL conical flask and 250 mL of distilled water was added. The conical flask was covered with aluminium foil and kept in a reciprocating shaker for 24 h for continuous agitation at 150 rpm for thorough mixing. Then, the extract was filtered by using muslin cloth followed by Whatman no 1 filter paper. The resultant solution was kept for the nanoparticle synthesis.- Synthesis of maize starch20 g of maize starch BP was dispersed in 50 mL of distilled water in a 250 mL beaker to provide a starch-water ratio of 0.40. The slurry was mixed for 60 min at 25°C to fully suspend the starch granules. After the pH was adjusted to 8.0 with 1 M aq.Sodium hydroxide (1M NaOH), 8% w/w acetic anhydride, was added in drops. 1M NaOH was added simultaneously at a rate of 20 mL/min to keep the pH of the suspension at 7.8 to 8.2 during the reaction. The reaction continued for I h at the completion of acetic anhydride incorporation. The slurry was adjusted to pH 5.5 with 1 M hydrochloric acid to end the reaction. The slurry was vacuum-filtered and the resulting cake mixed with 40 mL of deionised water and filtered again. The resultant cake was washed thoroughly thrice with deionised water to remove the remaining acid and then dried overnight in an oven at 40°C. The modified starch (MS) was stored for subsequent use.0.25, 0.5 and 0.75% modified Starch (MS) were prepared. This was done according to the method of Jackson et al., 2016 with slight modification by dispersing 0.25g, 0.5g and 0.75 g in 100ml of deionised water in a beaker maintained at 40°C in a water bath.- Synthesis of silver nanoparticles using aqueous extracts of Acalypha ciliata with model drugTo each of the starch dispersions previously prepared, 1% AgNO3 (10 mL) was added under constant stirring on a magnetic stirrer assembly for 5 min, followed by incorporation of Metronidazole (400 mg) to obtain [Ag (Drug/MS)] + dispersion.25 mL aliquot of a freshly prepared aqueous extract of Acalypha ciliata leaves (reducing agent), was introduced to the resultant mixture and maintained at 40°C temperature for 24 h. The resultant suspension of Ag/drug/starch was freeze dried (Gallenkamp, England) and subjected to further analysis.B. Characterisation of Silver nanocomposites- UV – vis spectroscopy to determine surface Plasmon resonance for silver nanoparticlesAnalysis of spectra was carried out with UV vis spectrophotometer (UNICO 2100, China). Scanning was done at a wavelength of 200 to 800 nm.Percent Yield, Entrapment Efficiency and Loading capacityThe percentage yield was calculated by dividing the actual yield by the initial weight of sample and multiplying the result by 100 percent.50 mg of the nanoparticle was dissolved in 50 ml of Phosphate buffer (pH 6.8). The suspension was ultra- centrifuged at 1500 rpm at 4°C for 30 min. The supernatant was analysed for metronidazole at 272 nm. The loading capacity and entrapment efficiency were determined for each nanoparticle using the formulae below:1. Loading Capacity (L.C) = (weight of Metronidazole in the nanoparticle – weight of Metronidazole in the supernatant) /weight of Metronidazole nanoparticle x 1002. Entrapment Efficiency (E.E) = (weight of Metronidazole in the nanoparticle – weight of Metronidazole in the supernatant)/weight of Metronidazole in the nanoparticle x 100- Determination of Particle Diameter and Index of polydispersity(PDI).The mean diameter (Z-average) and PDI of the nanoparticles were determined by dynamic light scattering method on a Zetasizer Ver: 7.01 (Nano Zs 90, Malvern Instruments Ltd, UK).Differential Scanning Calorimetry (DSC)Thermogram of drug- polymer was employed for the analysis. Scanning was done from temperature range of 25-250°C at the rate of 50°C/min.- Morphological Studies of nanoparticlesThe size and shape of nanoparticles were determined by Scanning Electron Microscope (SEM), Hitachi X650, Tokyo, Japan).- Release studiesRelease studies of Metronidazole in vitro were carried out separately in both simulated gastric fluid (pH 1.2 - acid buffer) and phosphate-buffered saline (PBS) solution (pH 7.4) using beaker magnetic stirrer assembly at 50 rpm at a temperature of 37±1°C. At interval of 30 min, 5 ml samples were collected and analysed at 272 nm for Metronidazole. 5 ml of fresh release medium was added to maintain sink conditions after each withdrawal.- Kinetic evaluationThe dissolution data of each batch were fitted to models like Zero order, First order and Higuchi to determine the release kinetics of the nanoparticles while the drug release mechanism was investigated by Korsmeyer-Peppas model.C. ANTIMICROBIAL STUDIES OF NANOPARTICLESThe MIC of optimized batch nanoparticles was determined.Microorganisms used:Four test micro organisms were used in this study; obtained from Pharmaceutical Microbiology laboratory in the Department of Pharmaceutics & Pharmaceutical Technology, University of Uyo, Nigeria. Gram-positive species used were Staphylococcus aureus and Streptococcus pyogenes while Escherichia coli and Psuedomonas aeruginosa, were the Gram-negative species. There were identified by their characteristics size and shape on specific media, followed by biochemical tests.Preparation of stock samples suspension:The nanoparticles were dissolved in distilled water to obtain a stock concentration of 10000µg/ml. The final strength of samples in the well was 5000 µg/ml. Metronidazole was prepared to a final strength of 16 µg/ml and served as the positive drug control for the bacterial strains.Preparation of innoculum:Inocula were prepared based on method recommended by CLSI [10]. Strains of bacteria were inoculated in Mueller Hinton agar and incubated at 35°C ± 2°C for 18 to 24 h. Suspensions of microbes in sterile saline solution were prepared from direct colonies. These suspensions were adjusted to a turbidity level of 0.5 McFarland approx. 1.5x108 CFU/mL [11].Determination of Minimum Inhibitory Concentration (MIC):The MIC of test pathogens and reference antibiotics were estimated by tetrazolium microplate assay [12].
3. Results and Discussions
Colour Change and UV – VIS spectroscopyFor NanoMet C, the Plasmon resonance at nanoparticle surface occurred at 407nm. The aqueous solution of silver nitrate changed from colourless to pale yellow on reaction with the Acalypha ciliata extract within 1 min and in 30 mins to yellowish brown indicating the presence of silver nanoparticles which was further confirmed by UV vis spectroscopy. The silver nanoparticles exhibited reddish brown colour due to excitation of plasmon resonance on particle surface. This colour change to dark-brown is as a result of increased strength and multiplication of silver nanoparticles. The UV-Vis spectrum of reaction medium displayed an emission peak at 407 nm, which corresponded to the absorbance of silver nanoparticles and showed well dispersed nanoparticles in the aqueous solution without aggregation in UV-Vis absorption spectrum.Table 1. Composition of Nanoparticles
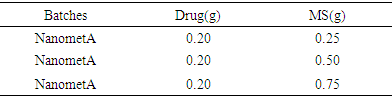 |
| |
|
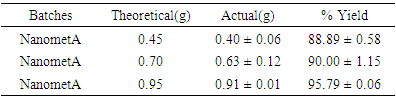
Percentage yield of nanocompositesThe percentage yields of the various nanoparticle formulations are presented in table 2. NanometA which consists of 0.25 %w/v Modified starch and 0.20 g metronidazole, had the least percentage yield (88.89) while NanometC consisting of 0.75 %w/v Modified starch and 0.20 g metronidazole had the highest yield (96.79%). The increase percentage yield may be due to increase in polymer concentration [13].Table 2. Percent yield of nanoparticles
 |
| |
|
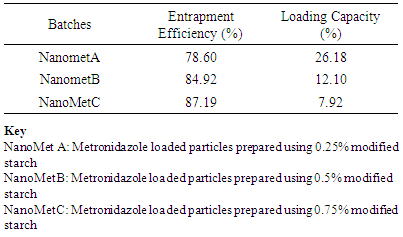
Entrapment Efficiency and Loading CapacityThe entrapment efficiencies and loading capacities of the nanoparticles are shown in Table 3. The entrapment efficiency was above 78% for all the formulations. Generally, there was significant (P < 0.05) increase in entrapment efficiency with increase in polymer concentration. The reverse was observed for the loading capacity; the increase in polymer concentration in the nanoparticles resulted in significant (P < 0.05) decrease in loading capacity. The highest entrapment efficiency was observed with NanometC (87.19%).Table 3. Entrapment Efficiency and Loading Capacity of Nanoparticles
 |
| |
|
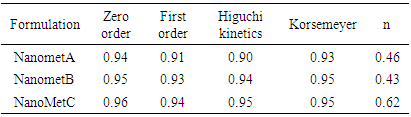
Drug loading expresses the percent weight of active ingredient encapsulated to the weight of nanoparticles. Drug loading efficiency is the ratio of the experimentally determined percentage of drug content compared with actual or theoretical mass of drug used for preparation of the nanoparticles [14]. The loading efficiency depends on the polymer–drug combination and the method used. Hydrophobic polymers encapsulate larger amounts of hydrophobic drugs, whereas hydrophilic polymers entrap greater amounts of more hydrophilic drugs [14]. Several formulation parameters, such as type of stabilizing agent and weight ratio of polymer to drug will influence the extent of drug loading [15,16]. Kinetics and Drug Release mechanismTable 4 shows the kinetics and mechanism of release of the formulations.Table 4. Kinetics and mechanism of release for Metronidazole nanoparticles
 |
| |
|
All the nanoparticles release the active drug via zero order kinetics (R2 = 0.94, 0.95 and 0.96 for NanometA, NanometB and NanometC respectively.Mean Particle sizeThe mean particle size of optimized nanoparticle, NanometC was 425,7 nm while the polydispersity index was 0.55. (table 5)Table 5. Mean particle size diameters and poly dispersity indices (PDI) of optimized nanoparticle
 |
| |
|
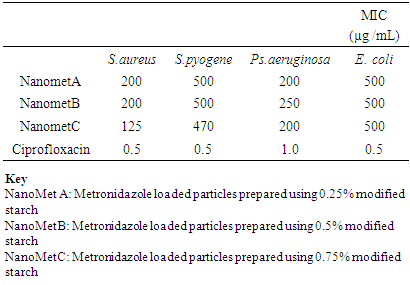
Antimicrobial StudiesThe nanoparticles were more effective against Staphylococcus aureus and Pseudomonas aeruginosa with MIC values of 125 ug/L to 250 ug/L.Table 6. Minimum inhibitory concentration (MIC) of nanoparticles
 |
| |
|
Differential Scanning Calorimetry (DSC)The thermograms of the optimized nanoparticle is shown in figure 1. DSC is used to obtain the thermal critical points like melting point, enthalpy specific heat or glass transition temperature of substances. It is a method of investigating the thermal characteristics of substances like polymers. DSC is a highly useful means of detecting drug-excipient incompatibility in a formulation. It gives insight into the capacity of the nanoparticles to entrap high amounts of the drug [17]. DSC detects phase transition such as glass transition (exothermic) crystallization and (endothermic) melting: the nanoparticle sample is heated and changes in the heat flow, compared to reference, are recorded [14]. DSC thermograms were obtained to define the physical state of the drug and polymer in the nanoparticles and to detect any drug polymer interactions in the polymeric network of the nanoparticles [14].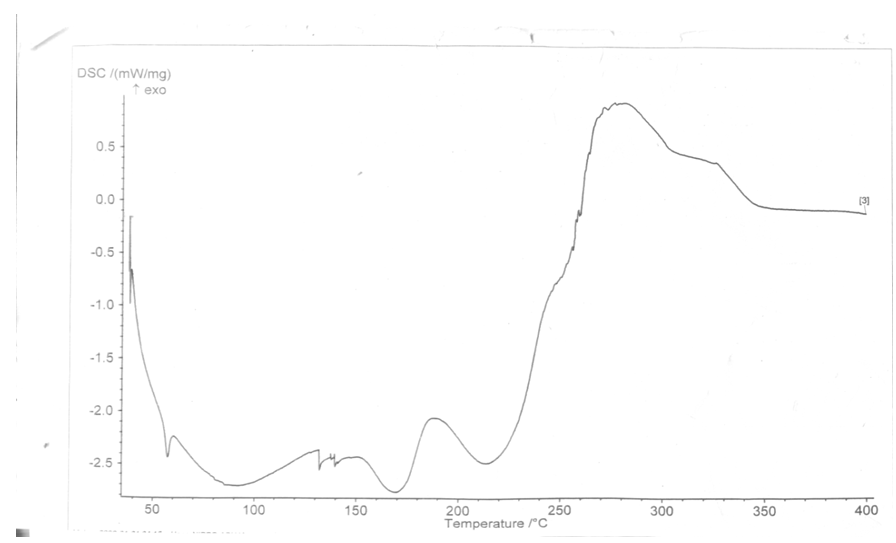 | Figure 1. Thermogram of NanometC |
Release Profiles of NanopartilcesThe maximum release of drug from Nanomet A (Fig 2) was 97 % at 8h. The time taken for 50% of the drug to be release was 6 h. The release of drug from Nanomet A followed a biphasic profile, with a slight initial burst at the first hour followed by sustained release for the next 8 h when the release study was performed in SGF. 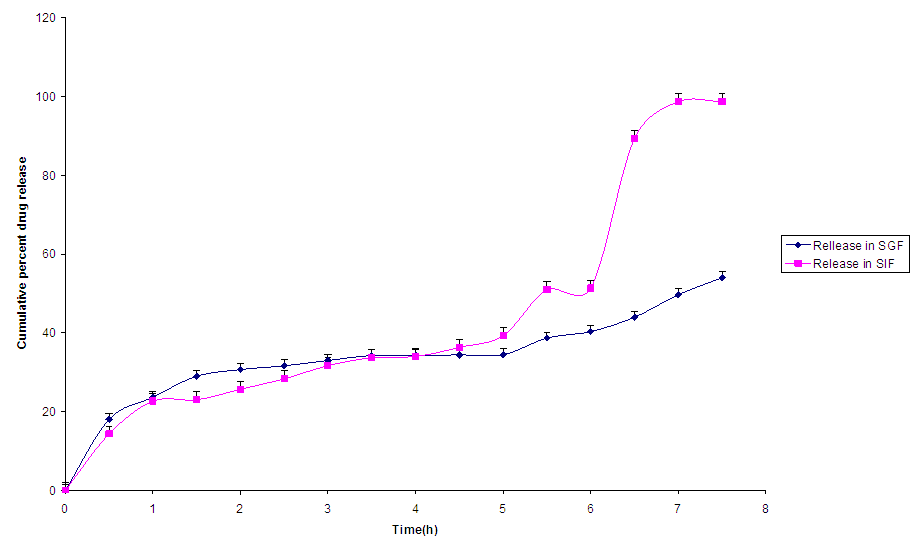 | Figure 2. Release profile of NanometA |
In SIF, the release of Metronidazole from NanometA showed triphasic profile, with a burst release within the first 1 h, followed by a lag phase between the first and second hour of release, after which, it was sustained till the 8th hour of study.The release profiles of NanometB (Fig. 3) followed a controlled release pattern. This may be due to diffusion of Metronidazole through the matrix coupled with polymer degradation. In SIF, the release was biphasic, with an initial burst release within the first hour and release was sustained till up to the 8th hour. 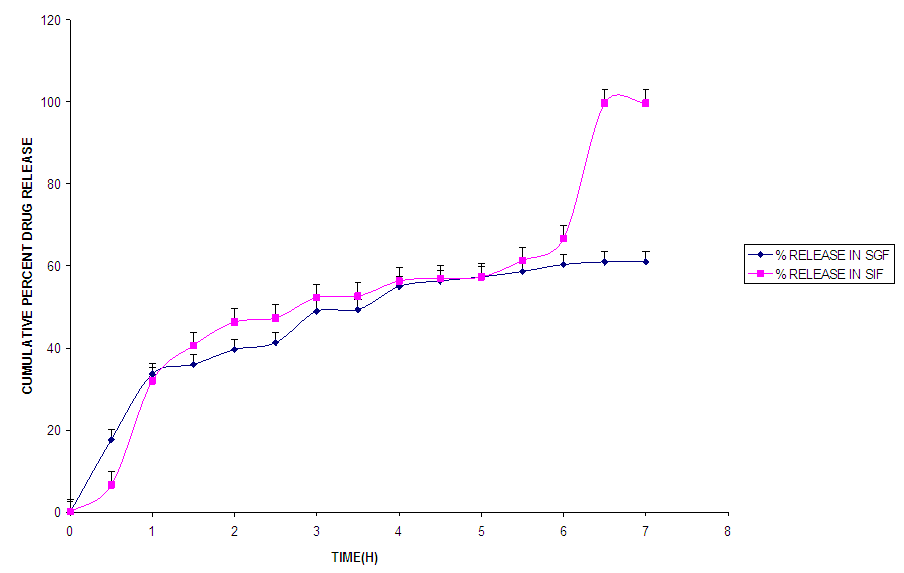 | Figure 3. Release profile of NanometB |
Nanomet C (Fig.4) showed a controlled release pattern in SGF but triphasic release profile in SIF. Release in both media followed zero order kinetics which involved polymer degradation and diffusion of the drug from the nanoparticles.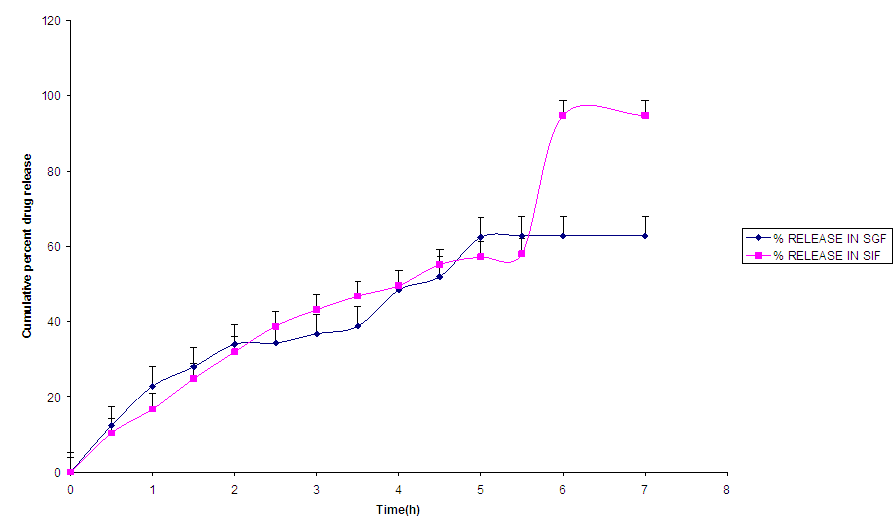 | Figure 4. Release profile of NanometC |
Kinetics and mechanism of release The kinetics and mechanism of release of the nanoparticles are presented in Table 4.NanometA formulation released the drug by zero order kinetics via non fickian diffusion (R2 = 0.94, n= 0.46), while the release of drug from NanometB followed zero order kinetics via fickian diffusion (R2 = 0.95, n= 0.43) [18]. The release of drug from this nanoparticle was independent of concentration. With further increase in polymer concentration, (NanometC), release kinetics followed zero via non fickian processes (R2 = 0.96, n= 0.62). NanometC released the drug through diffusion and erosion controlled mechanisms. The dosage forms following this profile, release the same amount of drug by unit time and it is the ideal method of drug release in order to achieve a prolonged therapeutic effect [19]).
4. Conclusions
In this research work, Metronidazole loaded silver nanoparticles were synthesised successfully with ecofriendly method using Acalypha ciliata as reducing agent and a semi- synthetic polymer as stabilizing agent. No record of this research finding is available in literature. The silver nanoparticles synthesis was confirmed through UV-visible spectrophotometer and characterized by scanning electron microscopy (SEM) and Dynamic light scattering (DLS). The nanoparticle synthesized was spherical to ovate in shape and were polydispersed in nature. The AgNPs showed good antibacterial activity against clinical pathogens and while in combination with commercial antibiotics (Amoxicillin and Carbenicillin) the antibacterial activity was further enhanced. Thus, we can conclude that the silver nanoparticles can be used as an antibacterial material alone and also in combination with the antibiotics but needs further studies to find out the mechanism behind the action of AgNPs on standard antibiotics and its enhanced bactericidal activity.UV VIS spectroscopy confirmed the presence of silver nanoparticles with absorption peak at 408.1nm. The particle diameter was 425.37 nm. The nanoparticles showed extended and controlled release profiles. The kinetics of release was predominantly zero order for the nanoparticles.The nanoparticles so formed had moderate antimicrobial properties.Nanoparticles prepared from semi synthetic starch (nanoMetC} can replace Metronidazole in the treatment of susceptible pathogens. The polymer, starch, used in this research, is biocompatible, affordable and readily available.
ACKNOWLEDGEMENTS
The authors are grateful to the staff members of Department of Pharmaceutics and Pharmaceutical Technology University of Uyo, Nigeria.
References
| [1] | Rao B, Tang RC. Green Synthesis of silver nanoparticles with antibacterial activities using aqueous Eriobotrya japonica leaf extract. Adv. Nat. Sci. Nanosci. Nanotechnol. 8. 2013; 015014 (8p). |
| [2] | Landage SM and Wasif AI. Nanosilver - an effective antimicrobial agent for finishing of textiles. International Journal of Engineering Science & Emerging Technology. 2012; 4(1): 66-78. |
| [3] | Korbekandi H, Ashari Z, Iravani S and Abbas S. Optimization of Biological Synthesis of Silver Nanoparticles using Fusarium oxysporum. Iran J Pharm Res. 2013; 12(3): 289 –298. |
| [4] | Mulongo G, Mbabazi J, Hak-Chol S. Synthesis and Characterisation of Silver Nanoparticles using High Electrical Charge Density and High Viscosity Organic Polymer Res: J. Chem. Sci. 2011; 1(4):18-21. |
| [5] | Rai MK., Deshmukh SD, Ingle AP, and Gade AK. Silver nanoparticles: The powerful nanoweapon against multidrug-resistant bacteria. J. Appl. Microbiol. 2012; 112, 841–852. |
| [6] | Stiufiuc R, Iacovita C, Lucaciu CM, Stiufiuc G, Dutu AG, Braescu C and Leopold N. SERS- active silver colloids prepared by reduction of silver nitrate with short-chain polyethylene glycol. Nanoscale Res. Lett. 8. 2013. doi:10.1186/1556-276X-8-47. |
| [7] | Jackson TC, Agboke A, Jackson I, Eti E. Biosynthesis of Silver nanoparticles using Murraya koenigii and acacia gum. Int. J. Res. Pharm.Sci, Bio, 2016; 3(2): 29 – 32. |
| [8] | Dos Santos CA, Seckler MM, Ingle AP, Gupta I, Galdiero S, Galdiero M et al. Silver nanoparticles: Therapeutical uses, toxicity, and safety issues. J. Pharm. Sci. 2014; 103: 1931–1944. |
| [9] | Rai M, Deshmukh SD, Ingle AP, Gupta IR, Galdiero M, Galdiero S. (2014). Metal nanoparticles: The protective nanoshield against virus infection. Crit. Rev. Microbiol. 2014; 1–11. |
| [10] | Clinical and Laboratory Standards Institute. Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria That Grow Aerobically; Approved Standard – Seventh Edition CLSI document M7-A7. Clinical and Laboratory Standards Institute. Wayne, Pennsylvania, United States. 2006. |
| [11] | Pamplona-Zomenhan, L.C., Pamplona, B. C., Barreto da Silva, C., Marcucci, M. C. Mimica, L.M.J. Evaluation of the in vitro antimicrobial activity of an ethanol extract of Brazilian classified Propolis on strains of Staphylococcus aureus Brazilian Journal of Microbiology 2011; 42: 1259-1264. |
| [12] | Shanmugapriya P, Suthagar P, Lee Wei C, Roziahanim M and Surash R. Determination of Minimum Inhibitory Concentration of Euphorbia hirta (L.) Extracts by Tetrazolium Microplate Assay. Journal of Natural Products 2012; 5:68-76. |
| [13] | Liu, D., Wu, Q., Chen, H., & Chang, P. R. Transitional properties of starch colloid with particle size reduction from microto NANO meter. Journal of Colloid and Interface Science, 2009; 339: 117–124. |
| [14] | Dhana lekshmi, U. M.; Poovia G and Neelakanta reddy. in-vitro observation of repaglinide engineered polymeric nanoparticles. Digest Journal of Nanomaterials and Biostructures 2012; 7(1): 1 – 18. |
| [15] | Dongming Peng, Kelong Huang, Yanfei Liu and Suqin Liu. Preparation of novel polymeric microspheres for controlled release of finasteride. International Journal of Pharmaceutics, 2007; 342 (1-2): 82-86. |
| [16] | Dhana Lekshmi, U. M; Poovi, G; Kishore, N and Reddy, N. In vitro characterization and in vivo toxicity studies of repaglinide loaded poly(methyl methacrylate) nanoparticles. International Journal of Pharmaceutics.2010; 396: 194-203. |
| [17] | Nnamani, P. O; Ibezim, E. C; Attama, A. A and Adikwu, M. U. Piroxicam- loaded peggylated tallow fat-based Solid lipid microparticles: characterization and in vivo evaluation. Nig.J.Pharm. Res 2010; 8 (1); 19 – 35. |
| [18] | Tahara, k; Yamamoto, K and Nishihata T. Overall mechanism behind matrix sustained release (SR) tablets prepared with hydroxypropyl methylcellulose 2910. J.Control Release, 1995; 35: 59- 66. |
| [19] | Chime Salome A; Onunkwo Godswill C. and Onyishi Ikechukwu I., Kinetics and Mechanism of Drug Release from Swellable and non Swellable matricies: A Review; Res.J.Pharm. Bio. Chem. Sci 2013; 4(2): 97 – 103. |






 Abstract
Abstract Reference
Reference Full-Text PDF
Full-Text PDF Full-text HTML
Full-text HTML