-
Paper Information
- Paper Submission
-
Journal Information
- About This Journal
- Editorial Board
- Current Issue
- Archive
- Author Guidelines
- Contact Us
Journal of Microbiology Research
p-ISSN: 2166-5885 e-ISSN: 2166-5931
2017; 7(4): 79-92
doi:10.5923/j.microbiology.20170704.01

Nitrogenase and Hydrogenase of the Actinobacteria Frankia: From Gene Expression to Proteins Function
Richau Kerstin H. 1, Pujic Petar 2, Normand Philippe 2, Sellstedt Anita 1
1Department of Plant Physiology, Umeå Plant Science Center, Umeå University, S-90187 Umeå, Sweden
2Université Lyon 1, Université de Lyon, CNRS, Ecologie Microbienne, UMR 5557, INRA 1418, Villeurbanne, Cedex, France
Correspondence to: Sellstedt Anita , Department of Plant Physiology, Umeå Plant Science Center, Umeå University, S-90187 Umeå, Sweden.
| Email: |  |
Copyright © 2017 Scientific & Academic Publishing. All Rights Reserved.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/

Frankia R43 is a filamentous actinobacteria, known for its ability to fix atmospheric nitrogen and to concomitantly produce hydrogen. Not much is understood about the variations in nitrogen fixation and hydrogen production among Frankia strains at the molecular level. Thus, studies providing direct comparisons between less known and well-annotated Frankia strains, may provide further information on the specific functions of the nitrogenase and hydrogenase(s). We compared candidate gene expression, protein sequence similarities and physiological activities of the well-annotated clade I Frankia casuarinae strain CcI3 and the recently sequenced lineage III Frankia strain R43. Our candidate genes, members of the nif gene cluster, involved in dinitrogen fixation, and genes encoding uptake hydrogenase (Hup) subunits, with an importance for energetic efficiency, were examined under varying oxygen and nitrogen conditions. Relative expression of almost all genes tested was higher in CcI3 than in R43, except nifK, following different but distinct expression patterns among all conditions. Candidate genes expression in CcI3 seemed more sensitive to oxygen than in R43, and the oxygen status seemed to influence gene expression much more than the nitrogen status per se.
Keywords: Actinobacteria, Frankia, Hydrogenase, nif -genes, Nitrogen-fixation, Gene expression
Cite this paper: Richau Kerstin H. , Pujic Petar , Normand Philippe , Sellstedt Anita , Nitrogenase and Hydrogenase of the Actinobacteria Frankia: From Gene Expression to Proteins Function, Journal of Microbiology Research, Vol. 7 No. 4, 2017, pp. 79-92. doi: 10.5923/j.microbiology.20170704.01.
Article Outline
1. Introduction
- Since the fixation of atmospheric nitrogen by the enzyme nitrogenase leads to the concomitant evolution of energy-rich free hydrogen (Schubert and Evans, 1976, Mohapatra et al., 2004), oxidizing the evolved hydrogen is an alternative way of generating energy. Bacteria capable of hydrogen production, besides that originating from nitrogenase, possess enzymes called hydrogenases that catalyze the simple reaction 2H+ + 2e-<-> H2 (Robson et al., 2001). Hydrogenases are extremely efficient catalysts for the production or oxidation of hydrogen, at rates exceeding thousands per second near the reversible potential, and establishing the mechanisms by which they do so is crucial for the development of future H2 technologies (Vignais and Billoud, 2007, Vincent et al., 2007, Armstrong et al., 20013). Frankia is known to oxidize H2 through the use of a [Ni-Fe] hydrogenase, which is coordinated by special ligands and buried inside the enzyme known as uptake hydrogenase (Hup) (Mattsson et al., 2001, Fontecilla-Champs et al., 2007, Richau et al., 2013). [Ni-Fe]-hydrogenases are the most extensively studied of the three main groups of hydrogenases identified up to date, and they also representing the largest group, consisting of 4 subgroups (Vignais and Billoud, 2007). Recently a model of [Ni-Fe]-hydrogenase has been developed that mimics the function of naturally-occurring [Ni-Fe]-hydrogenase (Ogo et al., 2013). In Frankia it has been shown that Hup consists of two subunits, a large one (HupL) and a small one (HupS) (Mattsson et al., 2001, Richau et al., 2013), and that Frankia alni strain ACN14a has two uptake hydrogenase syntons, one of which is expressed under free-living conditions and the other under symbiotic conditions, they are known as synton 1 and synton 2 respectively (Leul et al., 2007). Interestingly, Leul et al. (2007) showed that the small subunit from the first synton (HupS1) in Frankia ACN14a was more similar to HupS1 in Frankia CcI3 than to its own synton 2 small subunit (HupS2), which is evocative of an ancient duplication of the synton. In addition, Frankia strains have been identified to belong to 4 different lineages, with R43 being a member of clade3 while CcI3 is a member of clade I (Tisa et al., 2016). Frankia bacteria are heterotrophic actinobacteria, well known for their symbiotically facultative life-style and for their ability to fix nitrogen under both free living and symbiotic conditions (Tisa et al., 2016). Within the present work, we aim to replace the concept of the existence of “atypical” strains by introducing the term “proteosymbiotic” strains. The term “atypical” might lead to confusion, because in present literature, the term is used for either lineage IV strains with no nif genes and no symbiotic abilities, but the ability to sneak into datisca and coriaria or for strains clade III which have nif genes, are fully symbiotic on elaeagnaceae and have the ability to sneak into casuarina following clade Ic. To avoid any confusion we introduce the term “proteosymbiotic” for Frankia strain R43 because it has the two described symbiotic modes, has nif genes and is a member of the lineage III. R43 was originally isolated from Casuarina cunninghamiana Miq. (Berry et al., 2011), possesses specific characters, such as a) not being (re-) infective on its own host (Nod-) but able to infect the permissive Hippophae and Morella (Baker, 1987), b) being able to effectively fix nitrogen (Fix+) and c) to produce free hydrogen (Mohapatra et al., 2004). Hence R43 represents an interesting candidate for investigations at the molecular level in comparison with Frankia strain CcI3, which is well-annotated and completely sequenced (Lechevalier, 1986). CcI3 is considered a typical Frankia strain, since it is infective on its host plants and fixes nitrogen only under N starvation conditions. However, CcI3 is not known to produce free hydrogen other than that from nitrogenase.Successful efforts to sequence the genomes of Frankia strains ranging from Frankia ACN14a, EAN1pec, CcI3 (Normand et al., 2007), Candidatus Frankia datiscae Dg1 (Persson et al., 2011), Frankia CN3 (Ghodbane-Gtari et al., 2013) to Frankia BCU110501 (Wall et al., 2013), and our most recent R43 sequencing approach (Pujic et al., 2015) have greatly improved our knowledge of the genus. Interestingly, expression of genes necessary for ammonium assimilation is reduced when Frankia is in symbiosis, suggesting that symbiotic Frankia may not perceive N starvation and that the nitrogenase-encoding nif genes are regulated by signals other than those involved in N metabolism for instance by O2 level (Alloisio et al., 2010). Genes known to be involved in nitrogen fixation, encoding proteins such as Nif (nitrogenase) and Hup (uptake hydrogenase), are scattered throughout the genomes of Frankia strains (Lechevalier, 1986). Alloisio et al. (Alloisio et al., 2010) showed that nodule up-regulated genes in Frankia alni are mostly distributed over a number of regions that show high synteny between three Frankia genomes and that, as expected, several genes linked to nitrogen fixation were up-regulated in nodules. Amongst these were genes belonging to the nif gene cluster, a large gene cluster found in all nitrogen fixing bacteria, known to encode various enzymes involved in the fixation of atmospheric nitrogen. The enzyme encoded by the nifH gene is the nitrogenase reductase, which has a [Fe4-S4] cofactor and transfers electrons to the dinitrogenase (Hu and Ribbe, 2013). NifH functions as dinitrogenase reductase. More precisely, it transfers electrons gathered from ferredoxins to the two subunits of the dinitrogenase itself, coded by the nifD and nifK genes. NifD (α-subunit) and NifK (β-subunit) are the two subunits of dinitrogenase, forming a heterotetrameric enzyme with two FeMo-cofactors (Benson et al., 2011, Spatzal et al., 2011, Rubio and Ludden, 2008, Schwarz et al., 2009), which converts atmospheric nitrogen (N2) into ammonium and free hydrogen (H2). Additionally, another group of about 15 functional proteins is involved in protein folding and maturation as well as nitrogen fixation. Nif genes are present in all nitrogen-fixing organisms, as well as in nitrogen-fixing Frankia sp., and the latter expresses nif genes under both free-living conditions and when in symbiosis with various actinorhizal host plants. Their expression is believed to be induced in response to low concentrations of fixed nitrogen and low oxygen pressure within the nodule environment as well as those in soil under non-symbiotic conditions. The second group of genes up-regulated in nodules includes the hydrogen uptake hydrogenase (Hup), hup genes, which are known to be crucial for nitrogenase efficiency. In nitrogen-fixing organisms, hup genes play a key role in energy management by converting the hydrogen produced by nitrogenase activity into electrons and protons, thus, Hup proteins recycle energy for various other processes that would otherwise be wasted. Factors known to affect hydrogenase activity include O2 and H2 levels (Alloisio et al., 2010). As in Rhizobium leguminosarum, the expression of the hup genes in Frankia is concomitant with that of the nitrogenase-encoding genes (Lawson and Smith, 2002). Within the present work, we compared Frankia R43 and CcI3 with respect to genome size as well as the sizes of candidate genes with products involved in nitrogen fixation. Furthermore, we investigated the expression patterns of three nif genes (nifH, K, D) and both syntons of the uptake hydrogenase genes, hup1and hup2, each of which contains genes for small and large hydrogenase subunits. We also multi-aligned the sequences of nitrogenase and hydrogenase proteins of the two strains investigated and compared with other Frankia strains from which candidate protein sequences were available online. Based on these protein sequences, we constructed cladograms to gather more information about the relationship between the recently sequenced Frankia strain R43 and other Frankia strains.
2. Results
2.1. Genome and Gene Sizes Comparison
- There was a distinct difference in genome sizes between the two strains investigated in this study, with CcI3 having a genome consisting of 5.4 Mbp, in comparison to the R43 genome of 10.45 Mbp in size, a size difference of about 52% (1.9 fold). We found all candidate hup genes (hupL1, S1, L2, S2) present in Frankia R43 (Table 1). While the gene sizes of both subunits of synton 1 and the large subunit of the second synton do differ by about 120bp, the gene of the small subunit of the synton 2 hydrogenase of R43 differs by 309bp in comparison to CcI3. There were no significant differences in sizes of the nif genes found between the two strains. However, in both strains (=/- 3-6bp), aside from the nifD genes differing by about 200bp, with R43 possessing the bigger gene. However, in both strains under investigation, the nifH gene was just about 50% the length of nifD and nifK (Table 1). Therefore, all three nif proteins were similar in length in both strains, but the gene products encoded by nifD and nifK were about 1.7 and 1.8 times the size of that encoded by nifH, respectively (Table 1).
|
2.2. Gene Product Similarity Matrices
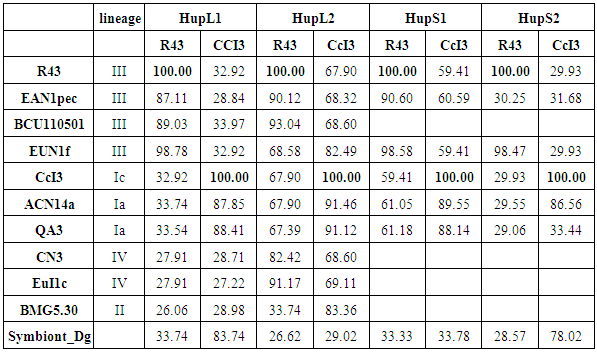
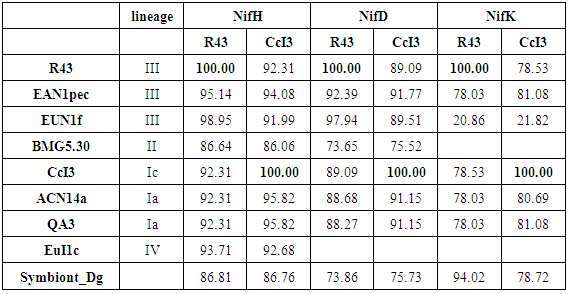
- Based on multiple alignments of our candidate proteins we established a matrix giving the percentage of similarity of candidate protein sequences (% similarity) among both strains under investigation and Frankia strains representing all existing lineages, for which candidate protein sequence information were available in public databases (Table 2). We also included the symbiont of Datisca glomerata (Symbiont_Dg).
|
|
|
2.3. Cladograms of Hup and Nif Protein Sequences
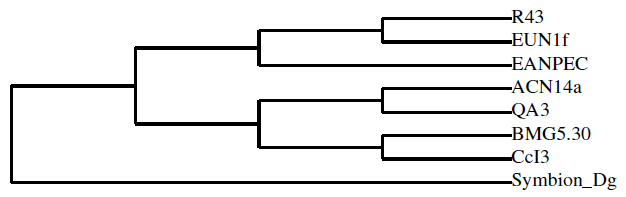
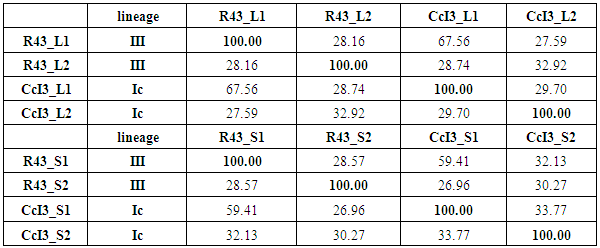
- Our cladograms (Figs. 1a and b), which were constructed based on observed distances between protein sequences, show that in the two strains the Hup protein sequences bore only weak similarities with one another, and the same is true for the Nif sequences. Moreover, the cladograms for the large Hup subunit of syntons 1 and 2 (Fig. 1a) clearly show that CcI3 and R43 are members of two distant clades. For both HupL1 and HupL2, the cladograms show that R43 clusters most closely together with Frankia EUN1f, also a strain with a large genome size(9.5Mb) compared to that of the CcI3 strain, whereas CcI3 clusters together with Frankia alni strain ACN14a, genome size 7.5Mb (16) and QA3, genome size 7.5 Mb (37). Our cladograms (Figure 1b) based on Nif protein sequences (NifH, D, K) similarly show a distant relationship between CcI3 and R43, with CcI3 showing high protein homology to the Frankia alni strain QA3 and R43 sharing a clade with EUN1f. Since our cladograms depicted in Figures 1 and 2 were established based on the multi-alignment files for the similarity matrixes above (Tables 2-4) they reflect these data. More precisely, the cladograms show that our candidate Hup as well as Nif proteins cluster specifically according to the lineages of their strains origin.
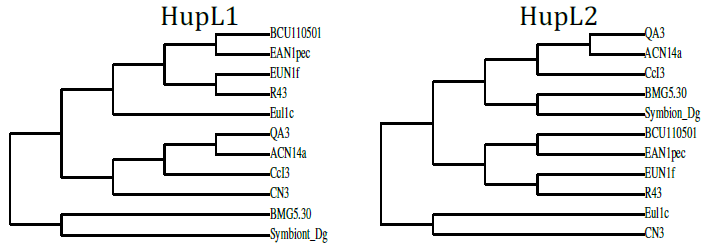 | Figure 1a. Cladograms based on observed distances between the large subunits of the two proteins HupL1 and HupL2 from various free living Frankia strains |
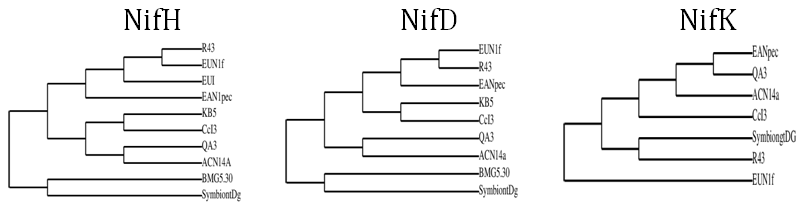 | Figure 1b. Cladograms based on observed distances between NifD, NifH and NifK protein sequences from various Frankia strains |
2.4. Candidate Gene Expression
2.4.1. Hup L1, S1, L2 and S2 Expression
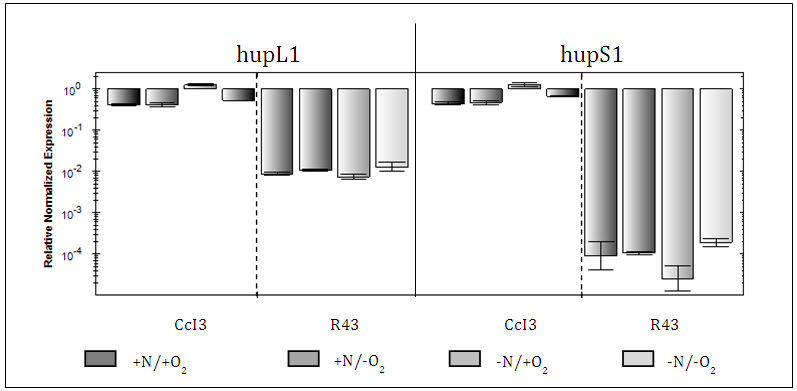
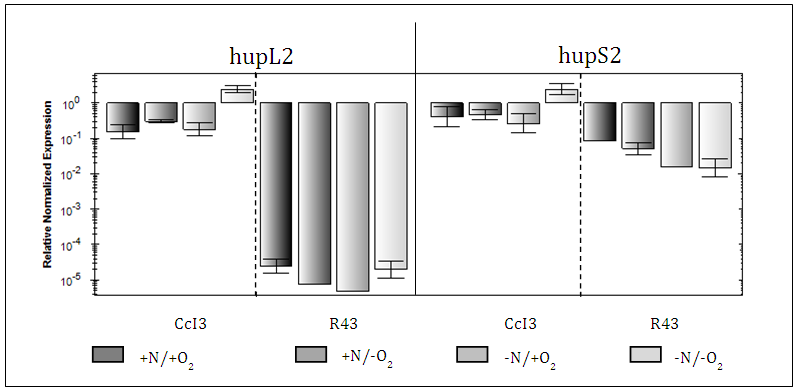
- Candidate genes expression was based on qRT-PCR experiments and their relative normalized expression was calculated based on the Frankia CcI3, 16S ribosomal RNA subunit as reference gene. Overall, Frankia CcI3 had hup expression levels much higher than that in R43 (>10000 times). Expression of CcI3 hupL1/S1 showed the highest expression under –N/+O compared to –N/ -O conditions, with an increase of about 2 times. Instead, hupL2/S2 expression peaked under -N/-O conditions followed by +N/-O conditions. The shift from deficient to sufficient nitrogen conditions made out an expression increase of about 10 time.R43, on the other hand, expressed all candidate hup genes at a much lower level regardless of the experimental conditions applied. In comparison to CcI3, R43 showed significant differences between the expression levels of hupL1/S1 and hupL2/S2. Frankia R43 Hup subunits encoded by the same synton, i.e. L1/S1 and L2/S2, were expressed independently of each other, in an unsynchronized manner. Expression of HupL1 in R43 was about 100 times higher than HupS1 expression, whereas R43 HupS2 was up to 1000 times more expressed than HupL2 in all treatments. Hup gene expression of R43 was shown to be highest under nitrogen sufficient conditions regardless of the oxygen conditions. In comparison, CcI3 demonstrated an increase of expression due to nitrogen deficiency.
2.4.2. nif H, D, K Gene Expression
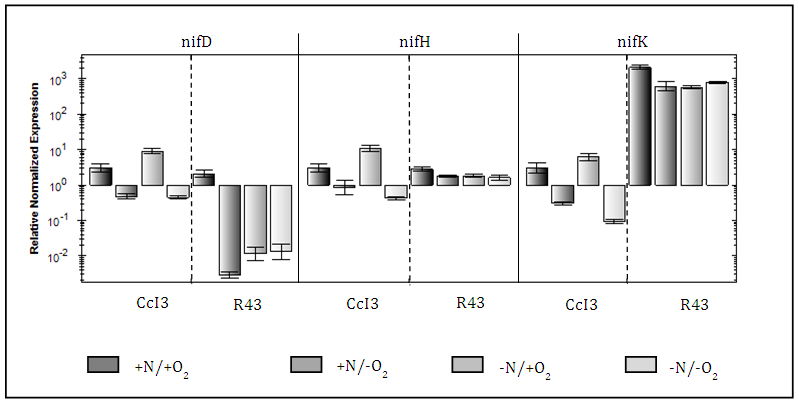
- Our expression analysis clearly shows that the patterns of expression differ significantly between strains for all nif genes investigated (Fig. 4). However, CcI3 shows distinct expression patterns for all three nif genes under the different oxygen and nitrogen regimes investigated. Fig. 4 shows that CcI3 expresses all nif genes at higher levels under anaerobic conditions than under aerobic conditions (3-10 fold differences). As compared to Frankia R43, CcI3 has overall higher relative normalized expression levels for the genes encoding NifH (up to 2-fold) and NifD (up to 4-fold), while in R43 NifK expression is up to 3.5-fold higher under all experimental conditions tested. Compared to Frankia CcI3, nif gene expression was found to vary significantly more overall in R43, except of nifK. Interestingly, expression of R43 nif genes does not follow any patterns induced by the experimental conditions applied. Neither +N/-O, -N/-O nor –N/+O conditions have significant effects on the expression of nif gene expression in this strain. However, +N/+O conditions increase the expression of nif gene subunits, especially that of nifD which increased 3.5 fold, while nifH gene expression was similar to that in CcI3 under all oxygen and nitrogen regimes. Compared to that of CcI3, the R43 nifK subunit is about 3 times more expressed, with the highest relative normalized expression being observed under +N/+O conditions.
2.5. Acetylene Reduction Assay (ARA)
- Over a timecourse of 12 days, we investigated nitrogenase activity in the two strains, namely R43 and CcI3, and a mixed culture containing both strains in equal densities. Figure 5 shows that the highest acetylene to ethylene reduction rate, thus activity of nitrogenase, was found for R43 at any time point measured. The mixed culture shows a slower onset of reduction than either strains grown singly, but on day 5 the mixed culture showed an equal activity as R43, meaning probably a C2H2 reduction rate higher than that of CcI3. Nevertheless, after 7 days the mixed culture activity declined and reached its peak below the reduction rate of CcI3 and R43 again. Since day 7 represents the peak of acetylene reduction capacity in all three cultures, we chose day 7 as the time point for harvest of bacterial cultures for subsequent experiments. At that time point R43 reduced about 26% and 35% more acetylene to ethylene in comparison to both CcI3 and the mixed culture, respectively.
3. Discussion
- As a symbiont, Frankia sp. thrives in association with a variety of dicotyledonous plants, so called actinorhizal species, from a range of habitats all over the world (Benson and Silvester, 1993). It has been suggested that Frankia strains may, as a consequence of adaptation to specific host plants, and/or geographic isolation of the host plant species, undergo genome shrinkage or expansion as they adapt to the new environments occupied (Baker, 1987, Normand et al., 2007, Persson et al., 2011). To be more specific, it has also been shown that genome sizes clearly correlate with the host range of the Frankia strains investigated, ranging from 10.45 Mb for Frankia R43 to 5.4 Mb for Frankia CcI3 and with the closeness of symbiotic association with smaller genome sizes seen for clade1c strain that are seen in soil outside their host plants (Normand et al., 2007, Pujic, et al., 2015).Moreover, the Frankia strain R43, a Fix+ but Nod- strain that cannot re-infect capacity to its original host plant Casuarina cunninghamiana after isolation, but having the ability to form nodules on other actinorhizal plant species such as Elaeagnus umbellata (. Benson and Silvester, 1993, Zhang et al., 1984) is belonging to lineage III. The genome of R43 is about 52% (1.9 fold) longer than that of CcI3 (Table 1) as shown in a recent publication (Pujic et al., 2015). Interestingly, Ghodbhane-Gtari et al. (2013) recently reported that Frankia CN3, another proteosymbiotic, Fix+ and Nod- strain has a genome size of 9.9 Mb (Ghodbane-Gtari et al., 2013), which might hint to the fact that clade 3 and IV strains in general have larger genomes than other Frankia strains.It has been shown and here confirmed that [Ni-Fe] uptake hydrogenases consist of large and small subunits that form heterodimers (Richau et al., 2013, Volbeda et al., 1995, Montet et al., 1997, Ogata et al., 2010), containing Ni and Fe atoms bound to the large subunit via four cysteine thiolate ligands in their catalytic center. Moreover, the small subunit contains either one [3Fe4S] or two [4Fe4S] iron–sulphur clusters that mediate electron transfer to and from the catalytic site (Volbeda et al., 1995). The importance of the four corresponding cysteine residues in [NiFe]-hydrogenases has been suggested to play a key role in maintaining the bacteriums’ oxygen tolerance of the cells (Liebgott, et al., 2011). Almost all the sizes of the candidate genes and their gene products presented here are comparable between CcI3 and R43, but interestingly not hupS2, which codes for the small subunit of the uptake hydrogenase expressed under symbiotic conditions. The small subunit of the uptake hydrogenase synton2 (HupS2) in the well-annotated Frankia strain CcI3 is approximately half the size of the HupS2 protein in Frankia strain R43 under investigation. The HupL2/S2 proteins appear to have diverged more in R43 than the HupL1/S1proteins which our cladograms reflect in Figure 1a. additionally, we suggest that the small subunit of the Hup protein in R43 may have an additional function beyond the mediation of electron transfer (Volbeda et al., 1995). Various functions of [NiFe] hydrogenases, including fermentation, energy conservation and hydrogen sensing, have been demonstrated in bacteria and archaea, depending on the enzyme's location and the type of iron–sulfur cluster, which are found in oxygen-tolerant membrane-bound [NiFe] hydrogenases (Fritsch et al., 2011, Laurinavichene et al., 2002, Shomura et al., 2011). We suggest that the larger size of the small subunit of the Hup protein in R43 may play a role in the higher oxygen tolerance of this strain, as evidenced by its expression patterns (Figure 1, 2). Moreover, since under free-living conditions the oxygen pressure to which the bacteria are exposed is much higher than that in symbiosis, it is likely that R43 gains extra protection against oxygen by means of expression of the larger HupS2 proteins from the Hup gene of synton 2, which shows a higher level of expression under free-living conditions (Leul et al., 2007). Actually, Leul et al. (2007) suggested that although the hydrogenase synton 1 and hydrogenase synton 2 code for uptake hydrogenase, but they may actually be coding for different types of uptake hydrogenases. One of the syntons is not simply a recent duplicate of the other, and functional complementarities can be expected. Given the obvious sequence divergence also shown in the present work we assume that the subunits as parts of the complete enzymes, might also be coded by different genes. Moreover, HupS1 and HupL1 distinctly diverged from HupS2 and HupL2 among the different Frankia lineages as seen in our similarity matrices Tables 2 and 3. These results indicated that hydrogenase synton 1 subunits of these Frankia strains were probably ancestral among the actinobacteria. Furthermore, Leul et al. (2007) assumed, since HupS2 and HupL2 of ACN14a and Frankia sp. CcI3, members of the lineage I, consistently grouped together and EAN1pec, member of lineage III was most closely related to the hydrogenases of non-Frankia bacteria, lateral gene transfer (LGT) between these organisms.Our similarity matrixes depicted two major results: a.) that the protein sequences bore a strong similarity within a given Frankia lineage and b.) that the first synton of the uptake hydrogenase seems to be more conserved that the 2nd synton, especially the large structural subunits. Since our cladograms depicted in Figure 1 and 2 were established based on the multi-alignment files for the similarity matrixes above (Tables 2-4) they reflect these data. More precisely, the cladograms show that our candidate Hup as well as Nif proteins cluster specifically according to the lineages of their strains origin.Our protein cladograms for the HupL subunit of syntons 2 and 3 show clearly that CcI3 and R43 are members of two distant clades. The cladograms for HupL1 and HupL2 are highly similar, clustering R43 most closely together with Frankia EUN1f, which is also a lineage III strain with a large genome (9.5Mb), whereas CcI3 clusters together with the well annotated Alnus- Frankia strains ACN14a, genome size 7.5Mb (Normand et al., 2007) and QA3, another strain with an intermediate genome size of 7.5Mb (Sen et al., 2013). Thus we checked the phylogeny of Hup and Nif proteins to compare with the16S-based one where R43 belongs to lineage III (large genomes, >9Mb), whereas CcI3 belongs to lineage Ic (small genomes, <6Mb), and ACN14a belongs to lineage Ia (intermediate genomes 7.5Mb) (Normand et al., 2007). Cladograms based on the observed distances between Nif protein sequences (NifD,H,K) similarly confirmed the distant relationship between CcI3 and R43, and CcI3 shows high protein homology to the other Alnus- Frankia strains (QA3), whereas R43 belongs to clade 3 together with EUN1f. According to our expression data presented here we suggest that in Frankia CcI3, hup genes expression of both syntons is constitutive but their expression levels alter when grown under different physiological conditions (Richau et al., 2013, Leul et al., 2007). Results in Figures 1 and 2 clearly indicate that CcI3 has levels of hup gene expression that are higher than that in R43, and that the relative level of gene expression in CcI3 does not significantly differ amongst the subunits or hydrogenase syntons. Hup gene expression in CcI3 exhibits a distinct pattern for both the small and the large subunit of synton 1 and 2, and CcI3 expresses all hup genes investigated most strongly under anaerobic, and particularly under –N, conditions, thus verifying CcI3 as a Nod+ nitrogen-fixing strain (Lechevalier, 1986), and hence hydrogen evolution during atmospheric nitrogen fixation in response to nitrogen starvation. Overall, candidate genes expression in CcI3 seems more sensitive to oxygen than in R43, and the oxygen status seems to influence gene expression much more than the nitrogen status per se. Moreover, the fact that the R43 hup genes expression is not altered by oxygen and the apparent oxygen sensitivity of CcI3 in comparison to R43, plus the fact that R43 encodes much larger hupS genes than CcI3, led to the supposition that the oxygen tolerant character of strain R43 is most likely to be based upon differences in enzyme architecture. This would be in line with the findings of Volbeda et al. (1995), who showed that [NiFe] hydrogenases may contain either one or two iron-sulphur clusters, ligands that are known to play a role in oxygen tolerance in bacteria and archaea (Fritsch et al., 2011, Laurinavichene et al., 2002).Overall, the miscellaneous annotations of the uptake hydrogenase gene as well as their gene products made a precise surveying of the structural subunits very challenging and therefore we suggest more regular data base updates and improvement in cross referencing of supplied data. For instance, single functional subunits of the hydrogenase were annotated with up to 7 different names and IDs.Although R43 expresses all candidate hup genes at a much lower level, in comparison to CcI3, regardless of the experimental conditions applied, the two subunits of the same gene seem to be expressed independently of each other, i.e. they are uncoordinated. In R43, expression of the gene encoding the small subunit of synton 1 (HupS1) is about 100 times lower than that of its large counterpart (HupL1), whereas in synton 2 the opposite situation is seen, the large subunit (HupL2) gene was much lower (about 100 times) expressed than the small one (HupS2) amongst all treatments. The fact that the small and the large subunit genes are expressed independently of each other under the experimental conditions used here suggests that the subunits of the Frankia R43 uptake hydrogenases may possess additional or altered functions either within or outside the Hup protein dimer, which is not documented up to date. Moreover, the difference in expression and in the size of HupS2 as well as the independent expression of the L and S subunits (Figure 2) lead us to the assumption that the small subunit of HupS2 has a special role in R43. In fact, there are several examples that microorganisms could have different types of hydrogenases, which would be expressed under different environmental conditions for specific purposes and with different roles (Richau et al., 2013, Shomura et al., 2011). Just as Leul et al. (2007) suggested that hydrogenase synton 1 and hydrogenase synton 2 may actually code hydrogenases genes for different conditions, we assume that the flexibility in an enzyme’s function might indeed be based on small differences in sequences of the subunit as parts of the complete enzyme and thus its function. With regards to our nif expression data, we were able to clearly show that the patterns of expression of all the nitrogenase genes/subunits that we investigated differed significantly between the two strains. Moreover, in contrast to CcI3, R43 showed little difference in expression of nitrogenase genes in response to different treatments. Fig. 1 shows that CcI3 expresses all nif genes at higher levels under anaerobic conditions than under aerobic conditions and + oxygen seemed to alter the pattern of nif genes expression pattern in CcI3 more than did nitrogen supply. Therefore, we conclude that CcI3 is more sensitive to + oxygen regimes than to nitrogen under the experimental conditions presented here. Additionally, it can be assumed that strain R43 is more oxygen tolerant, since a higher expression of all nif genes occurred under ambient oxygen concentrations. This higher tolerance of R43 to oxygen is correlated to the better adaptation to an autonomous life in the soil contrary to clade Ic strains such as CcI3 that are found only in intermediate proximity of their hosts.Expression of nifK gene in R43 is several times higher than that of the other nif genes, regardless of the experimental conditions applied. Moreover, R43 nifK was >1000 times higher expressed under +N/+O conditions than under any other conditions, possibly reflecting the atypical physiological characteristics of Frankia R43, and might hint again to much higher oxygen tolerance compared to CcI3. In case of elevated oxygen concentrations or enough reduced nitrogen the nifL subunit is activated which inhibits NifA, which activates all other nif genes. Therefore, NifL does not just act as feedback inhibitor for nitrogen oxidation but also as oxygen sensor. There is no nifL gene present in any Frankia strain sequenced up to date, which might indicate that the gene regulation as a function of oxygen or fixed nitrogen may occur based on other regulators.(Schmitz et al., 2002). It would be interesting to investigate nitrogenase regulatory circuits in R43, since the results might help to clarify the suggested higher oxygen tolerance of this strain. Frankia strain CcI3 and lineage Ic strains possess an oxyR-katG operon presumably involved in regulating peroxide stress, in other strains such as R43 (lineage III) or those of lineage I, the oxyR gene in a synton with a glutamate-cysteine synthase, that constitute the first step in the glutathione biosynthetic pathway and a glutamate-yielding asparagine synthase (Sen et al., 2013, Tavares et al., 2007). The independent expression patterns of the nif genes seems interesting for future investigations, since it is known that the genes of the nif gene cluster are part of the same operon.Within the ARA assay we used beside our two strains of investigation also an equal mix culture of both mixed culture), because attempted to see if both strains do interact just as they might do inside of nodules under symbiosis. We intended to answer the questions: 1.) Do our strains compete by means of acetylene reduction and/or 2.) Is R43 able to compensate for a lower activity of CcI3? The time course of ARA assay, a measure of nitrogenase activity, showed that R43 has an overall higher nitrogen fixing capacity at any time point measured, indicating a higher production of hydrogen under free living conditions. The fact that the mixed culture reduces its activity after 5 days to a level below that of the two candidate strains suggests an inhibiting effect induced by mixing the two strains under investigation. We suggest a competition between the two diverse strains and R43 seemed not have to be the ability to substitute for the lower reduction capacity of CcI3. However, that should be examined further in the future ie. by growth rate measures.
4. Materials and Methods
4.1. Frankia Strains and Culture Conditions
- The two Frankia strains under investigation, R43 and CcI3, were grown in continuous culture using the methods described by Mohapatra et al. (2004). In summary, bacterial cultures were initially grown in 50 ml of PUM medium, modified from P medium according to Shipton and Burggraaf (1982), at 27°C under constant shaking. After 7 days of incubation, half of each culture was transferred to PUM medium without N (-N) and the other half to PUM medium + N. All cultures were grown in glass vessels of capacity 100 ml, which were closed with rubber septa to prevent gas exchange with the ambient environment. Cultures purged with argon, referred to as anaerobic cultures, and cultures under -N conditions were supplemented with 10 mM NH4Cl to prevent hydrogen from being evolved by nitrogenase (Mohapatra, 2004). Both sets of bacterial cultures were incubated for an additional 7 days before they were again divided into two subcultures, of which one was purged with argon for 8 minutes, and the other was not purged to establish anaerobic (-O) and aerobic cultures (+O), respectively. Both anaerobic and aerobic cultures were then incubated for an additional 48 hours before RNA isolation was carried out for the expression analysis.
4.2. Genome Data Analysis
- The genomes of Frankia sp. CcI3, Frankia alni ACN14a and Frankia sp. EAN1pec were viewed using the MaGe tool in the MicroScope interface (Pruitt et al., 2007, Vallenet et al., 2006), and the Frankia EuI1c and Frankia EUN1f genomes were viewed on the DOE JGI homepage (Vallenet et al., 2013). Frankia R43 genes were identified after sequencing and assembling of the whole genome according to Pujic et al. (2015).
4.3. Multiple Alignments Proteins and Sequences Identity Matrix (% Similarity)
- Our aim was to align candidate protein sequences of Frankia strains representing all existing lineages (Table 2). We used the Frankia alni strain ACN14a (GenBank accession number NC_008278) as query for the blastp (protein-protein blast) suit on the NCBI homepage (Grigoriev et al., 2012, Altschul et al., 1997) and selected non-redundant proteins sequences and the Frankia taxid:1845 as organism as search set. Sequences of three members of lineage I were available and thus have been included into the multi-alignment studies of protein sequences. Frankia alni strain ACN14a (GenBank accession number NC_008278), the Casuarina-infective Frankia sp. strain CcI3 (NC_007777) and QA3 strains (CM001489.1). The sequencing of the lineage II strains, BMG5.30 is still in progress, but the partly available data were sufficient for or candidate protein alignment. Three clade 3- representatives have been sequenced, including Frankia sp. strains EAN1pec (NC_009921), EUN1f (NC_014666), EuI1c (ADGX00000000), BCU1105501 (ARDT00000000) and R43 (LFCW00000000.1), as well as clade IV strains CN3 and EuI1c with the accession numbers AGJN00000000 and CP002299.1 (12), respectively, as well as the symbiont of Datisca glomerata (Symbiont_Dg), member of the noncultured clade of Frankia strains, entering into root-nodule-symbioses with actinorhizal species from the orders Cucurbitales and Rosales (Persson et al., 2011). Protein sequences were assembled and transformed into fasta format as input for the Clustal Omega tool (Sievers et al., 2011, Goujon et al., 2010) a free online multi-alignment application.
4.3.1. Cladogram Construction
- We carried out so-called homology clustering and constructed cladograms based on the observed distances between Hup and Nif protein sequences from various strains representing all Frankia clusters together with our strains under investigation, R43 and CcI3, as well as the symbiont of Datisca glomerata. The Clustal Omega multi-alignment output (see above) was therefore up-loaded to the “One Click” method at Phylogey.fr homepage. TreeDyn (Dereeper et al., 2010, Dereeper et al., 2008, Chevenet et al., 2006), a tree editor, was used to create phylogenic trees, employing default workflow parameters.
4.4. RNA Extraction, cDNA Synthesis and qRT-PCR
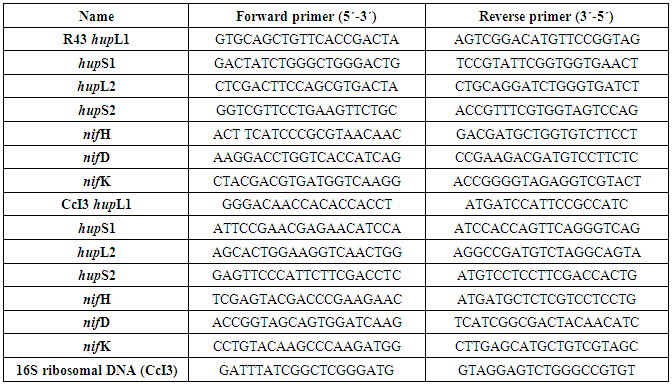
- All Frankia cultures were pre-treated according to the manufacturer’s protocol with RNAprotect (Qiagen, Hilden, Germany) prior to RNA isolation, to stabilize the RNA and eliminate all RNAses from the medium. After this pre-treatment, the bacterial cultures were shock-frozen in liquid nitrogen and the cell walls were disrupted by grinding with a mortar and pestle according to the RNAprotect Bacteria handbook. After mechanical disruption, we employed the RNeasy Plant Mini Kit (Qiagen, Hilden, Germany) to isolate RNA from the bacteria according to the manufacturer’s protocol. Immediately after isolation, the bacterial RNA was treated with RNAse-free DNAse (Invitrogen, CA, USA) for 30 min at 37°C, to remove contaminating DNA from the RNA samples. The integrity of the RNA was checked on a 1.5% agarose gel by visualizing the 16S and 23S RNA subunits under UV illumination. Additionally, RNA was loaded and analyzed on a Nano chip for the Agilent Bionanlyzer (Agilent Technologies, CA, US) and only RNA with a RNA integrity number (RIN) above 8,5 was used in our downstream applications.We used the iScript cDNA Synthesis Kit (Bio-Rad, CA USA), according to the manufacturer’s protocol to synthesize cDNA from 800-1000 ng of RNA. The amount of RNA was normalized based on Qubit™ fluorometric quantitation (ThermoFisher Sientific, Ma, USA). Subsequently primer efficiency was checked by carrying out PCR amplification using the applicable set of primers and template cDNA dilution series, and the optimal template/primer ratio was added to iQ SYBR Green Supermix (Bio-Rad, CA, USA) for the final qPCR analysis. Based on genome sequences obtained from the NCBI GenBank database (Pujic et al., 2015, Vallenet et al., 2013), 20 bp long qRT-PCR primers were designed with the help of the Primer3 program (Rozen and Skaletsky, 2000) using the default settings. Frankia CcI3 16S rRNA gene was used as a reference gene for the qPCR studies. Primer sequences for the candidate genes and the control Frankia 16S rRNA gene were listed within the appendix A, Table 1. qPCR was carried out on a CFX96 (Bio-Rad, CA USA), and the running conditions were as follows: 3 minutes at 95°C for initial denaturation and activation of the hot start polymerase in the reaction mix, followed by 40 cycles of denaturation (15 seconds at 95°C), annealing (30 seconds at 58°C) and extension (30 seconds at 72°C). Finally, a melting curve, from 55°C to 95°C with 0.5°C increments every 10 seconds, was prepared to detect possible contamination or primer-dimer formation within the assay mix.After qRT-PCR had run to completion, data analysis was carried out using the CFX Manager 3.0 software from Bio-Rad (CA, USA), with the parameters set according to Pfaffl (2001). Our relative normalized expression data are depicted relative to zero, with a log10 scale applied to the y-axes, and the error bars reflect the standard deviations of three technical replicates. Direct comparisons of candidate gene expression under nitrogen-replete and N-deficient, and under anaerobic and aerobic conditions, for both Frankia strains, R43 and CcI3, were carried out on the same plate.
4.5. Acetylene Reduction (ARA)
- Acetylene reduction as measure for nitrogenase activity was measured according to Mattsson and Sellstedt (2000). Therefore, we grew both strains as mentioned above and determined total protein concentration of both bacteria cultures via Bradford protein assay (Bio-Rad, CA, USA). The bacterial input was normalized, and a third mixed culture, consisting of both strains in equal concentration, was established with same protein input. Total protein input was about 400mg in all established 1ml sub-cultures, and the existing airspace in the culture vessel of same volume was replaced by 10% acetylene (C2H2). All cultures were incubated for 24 hour and acetylene reduction to ethylene was measured at day 2, 5, 7 and 9 after culture establishment according to former publication in the lab (Mattsson and Sellstedt, 2000). Acetylene reduction was established via gas chromatography (Shimadzu GC-8AIT, Shimadzu Scientific Instruments Inc., Colombia, MD) with ethylene as reference gas. All reference samples were to be found in the linear region of the established standard curve for a range of C2H4 concentrations.
ACKNOWLEDGMENTS
- J.C. Tryggers stiftelse CT12-435 (K.H.R., A.S.), Energimyndigheten 213-006259 (A.S.), Formas 942-205-92 (A.S.)
Abbreviations
- Hup: uptake hydrogenaseNif genes: genes involved in nitrogen fixation ARA: acetylene reduction assayNCBI: National Center for Biotechnology InformationDOE JGI: U.S. Department of Energy Joint Genome Institute
 Abstract
Abstract Reference
Reference Full-Text PDF
Full-Text PDF Full-text HTML
Full-text HTML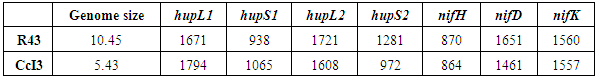
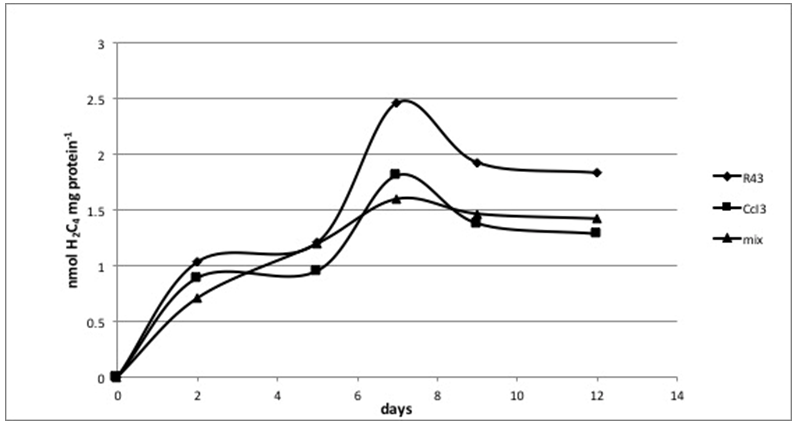
 C2H4) as measure of nitrogenase activity with 10% C2H2 as substrate for about 400mg of protein per sample (n= 8)
C2H4) as measure of nitrogenase activity with 10% C2H2 as substrate for about 400mg of protein per sample (n= 8)