-
Paper Information
- Previous Paper
- Paper Submission
-
Journal Information
- About This Journal
- Editorial Board
- Current Issue
- Archive
- Author Guidelines
- Contact Us
Journal of Microbiology Research
2012; 2(2): 19-25
doi: 10.5923/j.microbiology.20120202.04
Ribosomal Encoded Bacteriocins: Their Functional Insight and Applications
Jitendra Singh 1, Chaitali Ghosh 2
1School of Biotechnology, Gautam Buddha University, Greater Noida, 201308, Uttar Pradesh, India
2School of Biotechnology, Jawaharlal Nehru University, New Delhi, 110067, India
Correspondence to: Jitendra Singh , School of Biotechnology, Gautam Buddha University, Greater Noida, 201308, Uttar Pradesh, India.
| Email: |  |
Copyright © 2012 Scientific & Academic Publishing. All Rights Reserved.
Ribosomal encoded bacteriocins (25-80 kDa) are the secretory proteins which inhibit the growth of closely related bacteria. Their transcriptions are regulated under different environmental conditions and have different modes of action which includes non specific DNase activity, specific RNase activity, pore forming and inhibition of murein synthesis in bacteria. In this article we have summarized the genetic organization, regulation, structural organization, reception and functional activities of ribosomal encoded bacteriocins only and does not discuss peptide bacteriocins and microcins (<10 kDa). In the end of article practical applications of these bacteriocins has been described such as potential antitumor agent of their catalytic domain, biosensor for genotoxicity by exploiting their promoters and their effectiveness against HIV infection.
Keywords: Ribosomal encoded bacteriocins, nuclease, pore formation, biotechnological and therapeutic application
Article Outline
1. Introduction
- Bacteriocins are ribosomally encoded toxins produced by bacteria to inhibit the growth of similar or closely related bacterial strain(s)[1]. They are structurally, functionally, ecologically diverse and are produced by almost all the major lineages of Eubacteria and Archebacteria[2]. These ribosomally encoded bacteriocins are generally secreted in the extracellular medium by the producer where they recognize specific receptors on the surface of susceptible cells. They can induce toxicity in target cells by different mechanisms including enzymatic nuclease (DNase or RNase) as well as pore formation in the cytoplasmic membrane[3]. Their structure comprises of three distinct domain organization: (i) a domain involved in recognition of specific receptor, (ii) a domain involved in translocation, and (iii) a domain responsible for their toxic activity, with molecular mass of ~25 to 80 kDa[4]. Ribosomal encoded bacteriocins are broadly classified into two groups, groups A and B, based on cross-resistance. Group A comprises of bacteriocins that are translocated by the Tol system, such as colicins A, E1 to E9, K, L, N, S4, U, and Y, while group B comprises of bacteriocins that use the TonB system, such as colicins B, D, H, Ia, Ib, M, 5, and 10[3]. These proteins have received increasing attention for their potential use such as preservative in the food industries as well as in therapeutic applications for clinical usage. In this article we have focused on ribosomal encoded protein bacteriocins with their various aspects such as classification, genetic organization, regulation, secretion, reception, catalytic activities and their various applications.
2. Genetic Organization
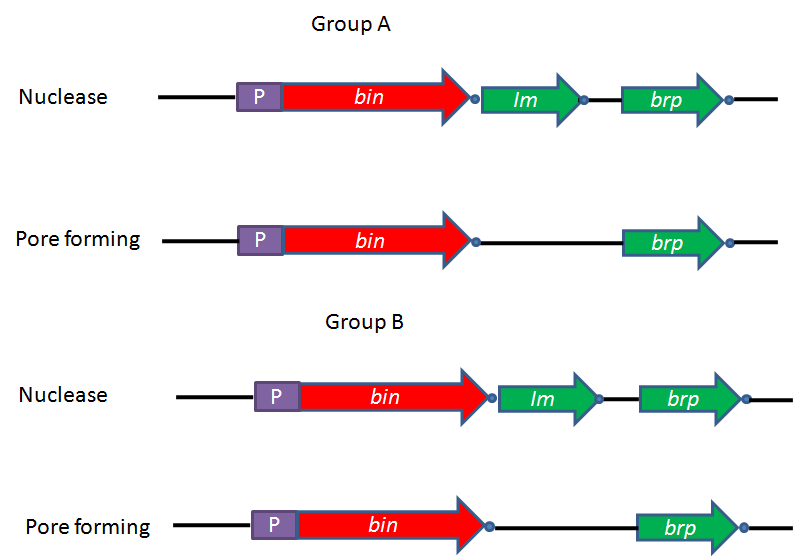
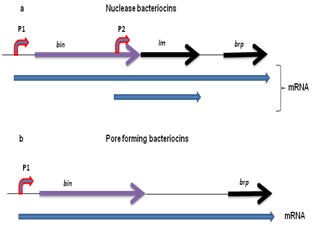
- The genetic organization of almost all known ribosomal encoded bacteriocins is summarized in Figure 1. In all the cases, the first ORF that encodes bacteriocin, called bin. It might be a unique gene in the case of operons of pore-forming bacteriocins of group B[5]. In the case of nuclease bacteriocins, gene encoding immunity protein designated as Im, generally located downstream of the structural gene for bacteriocin[6]. Immunity gene is under the regulation of two promoters: the LexA promoter of bacteriocin operon (P1 as shown in Figure 3 (a)) and its own constitutive operon (P2 as shown in Figure 3(a) which allows a constant production of the immunity protein in order to ensure that there is never any production of free bacteriocin, which could have detrimental effect on the producing cell. The consecutive promoter of immunity gene is located within the structural gene of the nuclease bacteriocin[7]. The operons encoding ionophoric bacteriocins contain no immunity gene; however it is located on the opposite DNA strand of the intergenic space between bacteriocin and lysis structural genes and is transcribed from its own promoter under constitutive regulation. The third gene of bacteriocin operon is the gene encoding for the lysis protein, which is named as brp (bacteriocin release protein). Product of this gene responsible for the release of bacteriocin into the medium as well as for the cell death after induction (colicins A, E1, E2, K, N, U, and Y and cloacin ). It is present in the operons of group A protein bacteriocins and in some operons of group B protein bacteriocins such as those of colicins 5[8], 10[9], and D[10]. Theses ribosomal encoded bacteriocin operons thus contain one to three genes.
3. Regulation
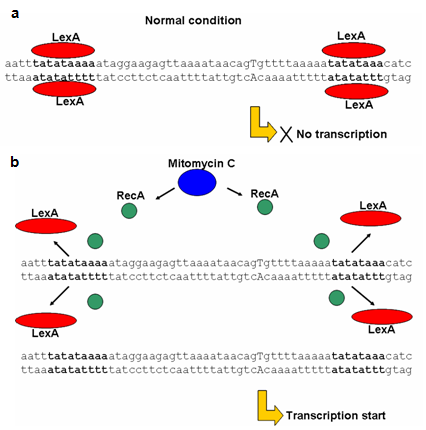
- Ribosomal encoded enzymatic bacteriocins are coordinately transcribed and translated with cognate immunity protein and after synthesis, form dimeric complex devoid of any enzymatic activity. Agents that are able to trigger the SOS response are also responsible for these bacteriocin productions they are of different nature like physical agents such as UV light to chemical drugs as well as stress conditions. Under normal conditions transcription of these bacteriocin operons is strongly repressed by the LexA protein, (the repressor of the SOS genes) except for cloacin DF13.
4. Structural Organization
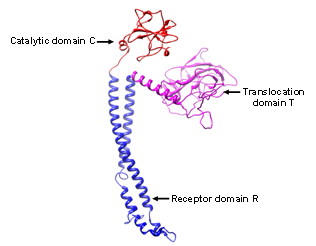
- Ribosomal encoded bacteriocins are generally organized into three domains performing three different functions including reception, translocation, and killing. The N’ terminal translocation domain (T) is important for translocation across the membrane of the target cell. Ton dependent bacteriocins contain a conserved five amino acid sequences called the ton box in their N’ terminus. Whereas Tol dependent bacteriocins have glycine rich and proline-rich region in this domain. The central domain is the receptor-binding domain (R) that binds to outer membrane receptors on the target cells e.g. colicin E9 binds to vitamin B12 receptor protein, Btu. The C-terminal domain comprises the cytotoxicity domain (C) which is pore-forming barrel-like domain in pore-formers and the DNase or RNase domain in the nucleases[3]. Protein bacteriocins are macromolecules containing an average of 500 to 600 residues, with a maximum of 697 residues for colicin D and a minimum of 271 residues for colicin M[26]. Crystal structure E3 colicin has been resolved which corroborate with the three domain organization of ribosomal encoded bacteriocin[27]. Three dimensional homology modeled structure of recently identified xenocin from X. nematophila containing translocation, reception and catalytic domain is represented in Figure 4 (unpublished data).
5. Reception
- Encounter between ribosomal encoded bacteriocin with its target bacteria is always occurs through the receptor domain of these bacteriocins. This is the first step in which these bacteriocins directly interact with the target bacteria. Outer membrane proteins of the target bacteria are the prime target of these bacteriocins[28]. In group A bacteriocins like colicins A and E1 to E9 target the TolB dependent vitamin B12 transporter BtuB. Colicin K binds to the nucleoside transporter Tsx, while colicin U interacts with OmpA. All of these bacteriocins need a second outer membrane protein for translocation (usually OmpF, but TolC is used by colicin E1), implying that initial binding interaction serves to position bacteriocin on the surface of target bacteria[29]. Therefore, primary receptor is not thought to participate in the translocation of bacteriocin across the outer membrane. All group A protein bacteriocins also require some combination of Tol proteins for translocation across the outer membrane. Group B protein bacteriocin use the Ton system for transit and most of them also bind to specific TonB-dependent transporters in the outer membrane. In these systems, bacteriocin appears to be bound and transported by a single outer membrane protein.
6. Activities
6.1. Nucleases
- Ribosomal encoded bacteriocins having nucleases activity fall into two groups, those which have nonspecific activity like DNase and those which have specific activity like RNase[30]. Most of these nuclease bacteriocins elicit cell death through either hydrolases or transferases by targeting phosphodiester bonds in the bacterial cytoplasm[31]. They target either genomic DNA (DNases), 16S rRNA (rRNases), or tRNAs (tRNases) and begin their passage into cells through either the Tol or Ton system[32-34]. After translocation of nuclease bacteriocin across the OM and periplasm, cytotoxic domain behaves differently from that of the pore-forming colicins, as the entire domain has to cross the cytoplasmic membrane. Their mechanism of translocation of nuclease domain into the cytosol must be independent of structure as they are structurally unrelated to each another [34]
6.2. Pore-Formation
- Pore forming protein bacteriocin kills sensitive bacteria forming ion channels in plasma membrane, resulting in membrane depolarization[35,36]. The structure of pore forming protein bacteriocin like Colicin A and four other pore domains have now been solved (of colicins Ia, E1, N, and B)[37-40]. These are compact water-soluble proteins and therefore, have ability to form voltage-gated ion channel in plasma membrane[41]
6.3. M type Protein Bacteriocin
- It is 28-kDa Ton-dependent ribosomal encoded bacteriocin, first described in the 1970s by Braun and colleagues[42]. It is one of the smallest ribosomal encoded bacteriocin known and not released in dimeric complex with its immunity protein. Its Immunity protein is located in the periplasm and anchored to the cytoplasmic membrane, where cytotoxic activity of the ColM is presumed to be expressed. Three dimensional structure of this immunity protein is recently deciphered by X-ray crystallography[43]. The ColM performed its cytotoxic activity by inhibiting both peptidoglycan biosynthesis and LPS O-antigen synthesis[26,44]. Recent study shows that cytotoxic effect of ColM occurs through the enzymatic degradation of ndecaprenyl phosphate-linked peptidoglycan precursors[45]. ColM target both lipid I and lipid II peptidoglycan intermediates and cleavage occurs between the pyrophosphoryl group and lipid moiety. Domain mapping and three dimensional crystal structure of ColM with crystallography has been recently studied[46,47]
7. Immunity Proteins
- Every nuclease bacteriocin is released from the producer cell in complex with an immunity protein, helping to prevent the death of producing organism. Nuclease-specific immunity proteins (Ims) are typically ~10 kDa and have ability to bind to the C-terminal nuclease domain to inactivate the protein bacteriocins by forming a ~71-kDa heterodimer. Protective mechanism for pore forming protein bacteriocin is achieved by a small polypeptide of 11 to 18 kDa, called the immunity protein, encoded by the same plasmid as bacteriocin. However, genome encoded immunity protein of xenocin from X. nematophila is having molecular weight of 42 kDa which is unique of its own[23] and has ability to form dimeric complex with xenocin (unpublished data). Using Immunity protein-specific antibodies Krone et al. showed that the Immunity protein for cloacin DF13 (a ColE3 rRNase homologue) is released at the cell surface[48]. It is shown that immunity proteins do not play a role in import of these bacteriocins as the DNase bacteriocins are fully active against E. coli regardless of whether the immunity protein is bound to bacteriocin or not[49]. From structural studies it has been shown that immunity proteins inactivate protein bacteriocin nucleases by one of the two distinct mechanisms: either by binding directly to the nuclease active site (tRNase- specific Im proteins) or by blocking the substrate binding site, leaving the active site open (DNase- and rRNase-specific Im proteins).
8. Sources
- Ribosomal encoded bacteriocins are generally produced by bacteria from Enterobacteriaceae family such as Enterobacter cloacae (cloacins), Yersinia pestis (pesticins), Klebsiella species (klebicins or klebocins)[50], Serratia marcescens (marcescins, colicins L and 24), Photorhabdus luminescens (lumicins), X. nematophila (xenocin)[23]. Escherichia coli (colicins)[51] and related species (Citrobacter freundii for colicin A or Shigella boydii for colicin U) are also known to produce bacteriocins to kill neighboring bacteria. Toxins with similar domain organizations and infection characteristics have been reported from Pseudomonas pyogenes (pyocins)[52], and even the gram-positive bacterium Bacillus megaterium (megacins)[53].
9. Applications of Bacteriocins
9.1. Biotechnological Applications
- Plasmid containment; in which origin of replication of plasmid ColE1 are often used to develop multiple copy plasmids which are stable and propagate in bacteria[54,55]. Biosensing of genotoxic compounds; the SOS regulated promoters of bacteriocins fused with lacZ gene can be used to test genotoxicity by calorimetry assay[56]. A system containing the SOS promoter/operator fused with GFP was recently developed to detect and quantify stress response induced by DNA-damaging agents[57,58]. Recombinant protein purification; bacteriocin released protein (BRP) have been exploited to induce cell lysis for recombinant protein release[59]. Bacteriocins are of interest in medicine because they are made by non-pathogenic bacteria that normally colonize in the human body. Loss of these harmless bacteria following antibiotic use may allow opportunistic pathogenic bacteria to invade the human body. Some bacteriocins are used commercially also, e.g. to prolong the shelf life of foods[60,61]. Effectiveness of the natural products like these ribososmal enocoded bacteriocins rather than chemical products give us further opportunity to use them in the control of plant disease of bacterial origin.
9.2. Therapeutic Application
- For therapeutic applications, E. coli strains producing bacteriocins are available under the names of MUTAFLOR and SYMBIOFLOR proved successful in the treatment of odiopathic chronic constipation as well as functional intestinal disorder and chronic inflammatory bowel disease[4]. Studies have shown that protein bacteriocin like colicins specifically inhibit the proliferation of cancer cells however, bacteriocin specificity for tumor cells is dependent on the cell line used[62]. It is observed that these ribosomal encoded bacteriocins are more toxic to tumor cells than to normal cells within the body[63] and moreover it did not show any toxic effect to the laboratory animals tested[64]. Studies have shown that these bacteriocins have ability to select cancerous cells from the normal cells but it is still unclear how this is accomplished. The eukaryotic cells naturally lack protein which inhibits protein bacteriocin. It could be possible that some of the natural inhibitors of eukaryotic cells such as RNase inhibitor protein that can bind variety of RNase, can bind also to certain bacterocins. Using bacteriocins like E3 colicin, it has been shown that eukaryotics cells are unlikely to contain an inhibitor protein specific for the rRNase. Therefore, it is unlikely that eukaryotic cells have any mechanism to inhibit the rRNase activity if the rRNase domain enters the cells[65]. Since, these bacteriocins are foreign protein to the patient, so it might be possible that immune system would attack and destroy them. However, the primary studies with mice showed that no activation of immune system[66,67]. In the long quest for medical applications, these ribosomal encoded bacteriocins have also been tested as AIDS drugs[68]. It has been studied that when bacteriocin is introduced to the AIDS lymphocytes infected with AIDS virus then uptake of the radioactivity is significantly reduced in 3 H-thymidine uptake test. While the growth of normal lymphocytes subjected to the same concentrations of bacteriocin was not impaired (71%). Interestingly, 57% of those normal lymphocytes which were subjected to protein bacteriocin showed an increase in radioactivity intake by comparison to those untreated. This signifies stimulation of growth of normal lymphocytes, a response which will presumably allow a substantial number of patients treated with protein bacteriocin greater immunity against other infections which attend AIDS patients. Recently Ogawa et al. have shown that Colicin E5 ribonuclease domain cleaves Saccharomyces cerevisiae tRNAs leading to impairment of the cell growth [69]. These kinds of studies will open new door for the therapeutic application of ribosomal endcoded bacterioicn against mycological diseases. Therefore, this broad range of applications of ribosomal encoded bacteriocins is certainly not exhaustive and may increase in the future.
10. Conclusions
- In this review we have describe how ribosomal encoded bacteriocins are genetically organized. Focusing on their regulation, under normal conditions transcription of protein bacteriocin gene is tightly regulated by LexA repressor protein. However, under stressful conditions, the SOS response turned on, and bacteriocin as well as its cognate immunity protein are co-expressed and secreted from the host cell as hetreodimeric complex. Structural studies revealed the domain organization of these bacteriocins which includes translocation domain, receptor domain and catalytic domain. Interaction with the target cells occur with receptor domain of bacteriocin and translocation of its catalytic domain into cytoplasm of the target cell occurs with assistant of either Tol or Ton proteins present in the periplasm of the target cells. Inside the cytoplasm of target cell, catalytic domain of these bacteriocin inhibits its growth by cleaving DNA nonspecifically or by inhibiting protein synthesis by specific cleavage of RNA. Pore forming bacteriocins depolarize target cell membrane and M type inhibits the synthesis of lipid I and lipid II peptidoglycan, and both the activities are responsible for the bacterial growth inhibition. From application point of view these ribosomal encoded bacteriocins are used as food preservative, biosensor and in medical usage. Property like anti-cancer makes these bacteriocins a good candidate as an alternative to conventional antitumor therapeutic drugs. In conclusion, these ribosomal encoded bacteriocins hold out the promises of better therapeutic as well as biotechnology applications and therefore, their studies in the future will certainly continue.
 Abstract
Abstract Reference
Reference Full-Text PDF
Full-Text PDF Full-Text HTML
Full-Text HTML