-
Paper Information
- Next Paper
- Previous Paper
- Paper Submission
-
Journal Information
- About This Journal
- Editorial Board
- Current Issue
- Archive
- Author Guidelines
- Contact Us
International Journal of Materials and Chemistry
p-ISSN: 2166-5346 e-ISSN: 2166-5354
2012; 2(2): 51-56
doi: 10.5923/j.ijmc.20120202.02
Rapid Mapping of Potential Active Site of Protein KSI by Multi-Dimensional NMR: Use of HyTEMPO as an Indirect Probe
Vandana Mehrotra1, Yong Nam Joe1, Hyung Jin Cha2, HyungJu Lee2, Kwan Yong Choi2, Hee Cheon Lee1
1Department of Chemistry
2Department of life Science Pohang University of Science and Technology, San 31 Hyojadong, Pohang, Pin code 790-784, Republic of Korea
Correspondence to: Vandana Mehrotra, Department of Chemistry.
| Email: |  |
Copyright © 2012 Scientific & Academic Publishing. All Rights Reserved.
A novel NMR method to rapidly map the enzyme -substrate active sites has been described. Potential active site mapping has been done by competition binding experiments using paramagnetic reagents (HyTEMPO) along with a substrate (or analog) (Equilenin), by 1H-15N HSQC NMR experiments. HyTEMPO induces the line broadening of signals in NMR spectrum. We analyzed the peak height value of cross peaks in 1H-15N HSQC spectrum of KSI with HyTEMPO and Equilenin. Our result revealed that residues in hydrophobic cavity of KSI, particularly active site region, were mainly perturbed by HyTEMPO. Upon the addition of Equilenin as an intermediate analog, few resonances were experienced as signal enhancements. These observations proved that HyTEMPO could map the potential active site of a protein KSI in the present case. This mapping method provides both high accuracy and speed, besides also a possibility of functional studies for unknown proteins in order to solve their difficult three-dimensional crystal structure.
Keywords: Hytempo, Equilenin, NMR Spectrum, Mapping, KSI Protein
Article Outline
1. Introduction
- The protein active site plays a significant role in various physiological and pathological processes in modern day biochemistry. The catalytic reaction is a binding reaction of protein similar to a lock and key model where the substrate behaves like a key which fits in to the lock symbolic to an active site of enzyme/protein by interaction between substrate and residues of active site in the protein. Depending upon the substrate conformation the active protein site fits to the of size, shape, charge, and hydrophobic or hydrophilic character similar to each other, or to be induced conformational changes in order to make fit binding to substrate.[1,2] Thus, the knowledge of the active site on protein is very important to understand the functions in proteins. Computational methods were mainly utilized for the earliest studies of active site prediction in proteins.[3-5], wherein the accuracies were low or not demonstrated experimentally. Thus, mapping of active site in protein by an experimental method is essential to understand the functions of proteins. Study of Enzyme mechanism carried out by several methods like X-ray crystallography, Nuclear Magnetic Resonance methods, florescence, and IR spectroscopy form an important part of structural biology.[6-8] Among these, X-ray crystallography and NMR are powerful enough to determine three dimensional structure of various molecules, inorganic compounds, DNA and proteins.[9] Moreover, these methods give accurate information for binding enzyme with the substrate. Because these two methods generally need much time and efforts to obtain structural information, to understand functions of unknown proteins. For this reason, the development of rapid mapping method for active site determination of reaction between substrate and enzyme becomes vital. 4 - Hydroxyl - 2, 2, 6, 6 - tetramethylpiperidinyl - 1 - oxy (HyTEMPO) is a nitroxide-soluble spin label that identifies the residues protected from the solvent. It has been used to distinguish solvent exposed area of the protein using multi dimensional NMR experiments.[10,11] Since the binding site between enzyme and substrate is mainly placed in hydrophobic core of most proteins, we hypothesize that it can be easily detected by NMR experiments using HyTEMPO in the present case..[12,13] Δ5-3-ketosteroid isomerase (KSI; EC 5.3.3.1) from Pseudomonas putida is a homo-dimeric enzyme which catalyzes the proton position conversion of 4β- to 6β.[14] It has been reported to contain hydrophobic residues at solvent exposed sites and has been studied extensively to understand the catalytic mechanism of the allylic rearrangement.[15,16] The structure of KSI using X-ray crystallography and NMR has revealed that this protein folds into three α-Helices and six β-Strands in each monomer.[6,17] In the present studies, the potential binding site of KSI was studied qualitatively by NMR experiment using HyTEMPO as an indirect probe. The potential binding site of KSI was very similar to the binding site of KSI as described by the crystal structure. These observations were confirmed by the cumulative chemical shift data.
2. Materials and Methods
2.1. Material
- 15N - labeled KSI Sample[18], 4 - hydroxyl - 2, 2, 6, 6 - tetramethyl-piperidine-1-oxy (HyTEMPO), deuterium oxide (D2O) and dimethyl sulfoxide (DMSO) were purchased from Sigma-Aldrich Company, USA (St. Louis, MO, 99.9 % purity). 15N-labeled NH4Cl were purchased from Cambridge Isotope Laboratories, Inc. (Andover, MA). These chemicals were used without further purifications. The intermediate analog, 3 - hydroxyl - 1,3,5(10),6,8 - estrapentaen - 17 - one (Equilenin), was purchased from Steraloids (Newport, RI, USA). It were dissolved in DMSO for the use of experiments and stored at room temperature.
2.2. Preparation of NMR Sample
- NMR samples (~1.5 mM) used for 1H-15N HSQC NMR spectra were prepared in a buffer (4℃) containing 20 mM potassium phosphate (pH 7.0, 10% v/v D2O), 1 mM DTT and a small amount of sodium 2,2-dimethyl-2- silapentane-5-sulfonate (DSS) for internal chemical shift reference of 1H and 15N. 4.5 mM HyTEMPO were added to the native state 15N KSI solution in order to analyze the binding region between KSI and substrate. A slightly excess amount of Equilenin relative to intermediate analog of the KSI was added to the 15N KSI solution with HyTEMPO. All NMR samples were equilibrated at room temperature for 48 hours after manipulations.
2.3. NMR Studies
- NMR experiment for backbone assignment of KSI- equilenin complex were performed on a Bruker 800 MHz spectrometer operating at 800.25MHz for proton frequency (Korea Basic Science Institute at Ochang) equipped with a triple resonance probe and pulsed field x-, y-, z- gradient capabilities. NMR spectra for backbone assignment of KSI-Equilenin complex were obtained by HNCA and HNCACB method. All 1H-15N HSQC NMR experiments were collected at 298K on a Bruker DRX-500 spectrometer operating at 500.13 MHz for proton frequency equipped with triple resonance probe. 1H-15N HSQC NMR spectra of NMR sample was recorded using a 90° pulse of 8.8 μs, 128 data points along t1 domain and 2048 data points along t2 domain using State-TPPI method in t1 dimension with a relaxation delay of 2 s.[19] Pulse sequence was modified to include gradient pulses and WATERGATE sequence in order to minimize artifacts and to avoid saturation of water resonances. The 1H chemical shifts were calibrated to DSS, and 15N chemical shifts were determined by indirect referencing.[20]
2.4. NMR Data Processing
- 1H-15N HSQC NMR spectra of native KSI, KSI with HyTEMPO and KSI with both HyTEMPO and Equilenin were recorded at the same condition except number of scan and receiver power. The NMR data were processed with an SGI octane 2 workstation using the NMRPipe software package.[21] All NMR data were analyzed using the Sparky software.[22]
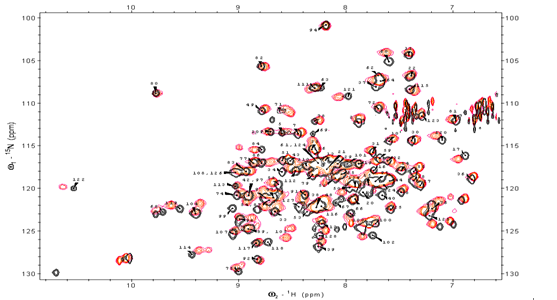 | Figure 1. 1H-15N HSQC NMR spectra of KSI |
3. Results
3.1. Resonance Assignment of KSI-Equilenin Complex
- Amide and 15N chemical shifts are sensitive probes for a given residue environment in a protein structure. The backbone resonance assignment of free-KSI was reported earlier and the number of peaks respectively present in the HSQC spectra is 120. The specific sequence of backbone assignments in steroid-bound KSI complex were primarily obtained by HNCA method. Among 131 residues in KSI, 110 residues except prolines and both terminal residues were represented in 1H-15N HSQC NMR spectra, and 96 residues were unassigned due to a severe spectral overlap (Fig. 1).In order to further investigate the local structural perturbation of proteins by Equilenin, cumulative chemical shift (Δδcum) was calculated for each residue according to the following equation.[26]
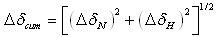 | (1) |
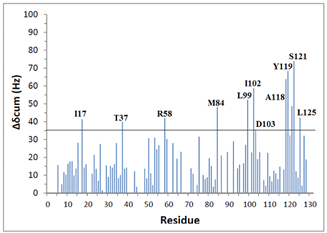 | Figure 2. Standard deviation values of some residues in KSI |
3.2. The effect of Paramagnetic Reagent HyTEMPO on Native KSI
- 1H-15N HSQC NMR experiments were performed to analyze paramagnetic effects, since two dimensional experiments provide sufficient resolution for the analysis of many residues. Fig. 3(a) shows finger print 1H-15N HSQC spectra of KSI samples with and without HyTEMPO. The peak height (intensity) of all of residues changed in 1H-15N HSQC spectrum. Specifically, for some residues it was particularly decreased, while some other residues disappeared in presence of HyTEMPO.which indicates that it causes a line-broadening for NMR signals of KSI residues. A detailed evaluation of paramagnetic effects, was done by calculating attenuations (Ai) of peak height by the chemical reagent according to the formula.
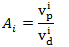 | (1) |
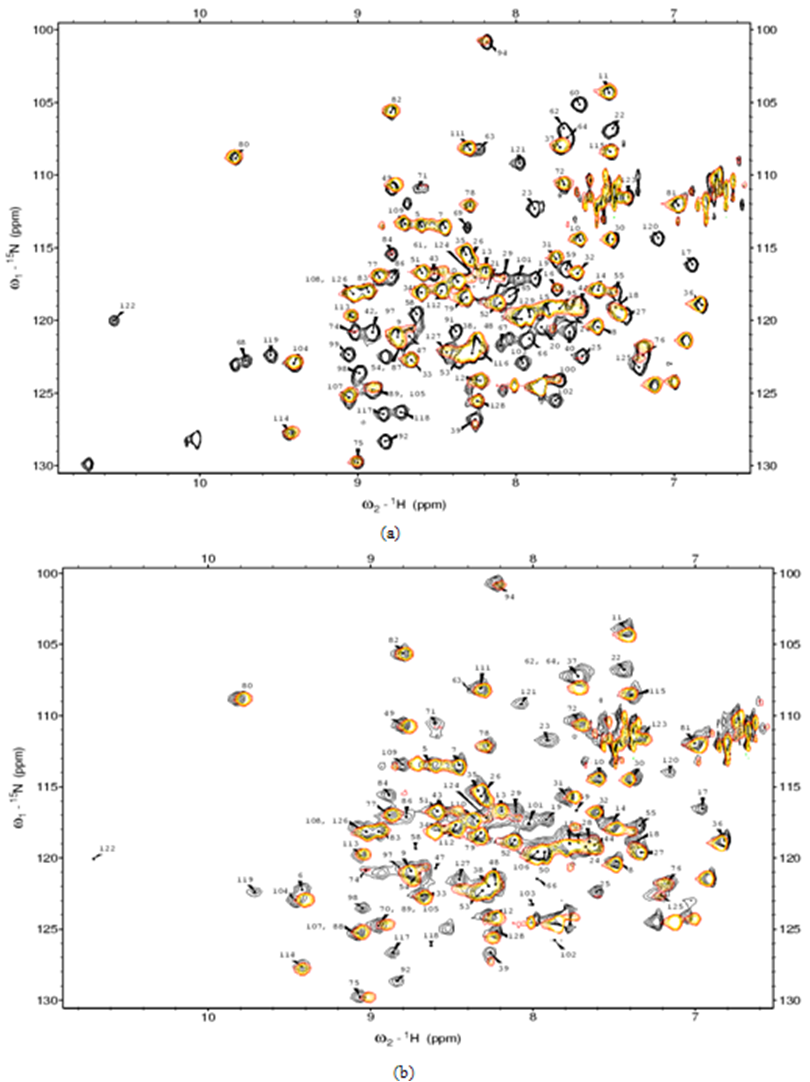 | Figure 3. Finger print 1H-15N HSQC spectra of KSI samples with and without HyTEMPO |
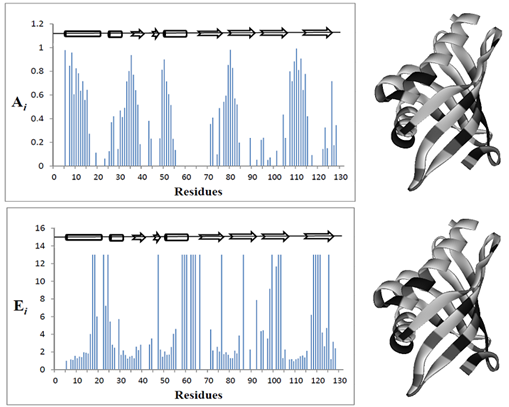 | Figure 4. shows Ai values as a function of amino acid residue number of KSI |
3.3. The NMR Signal Enhancements by Equilenin; the Intermediate Analog
- 1H-15N HSQC experiments were performed by monitoring the peak height in order to evaluate the effect of substrate on solution of KSI-HyTEMPO. Fig. 3(a) shows finger print 1H-15N HSQC spectra of KSI-HyTEMPO solution with and without Equilenin. As can be seen in Fig. 3, 1H-15N HSQC spectrum of KSI with HyTEMPO and Equilenin solution is almost identical to that of native KSI with Equilenin. Specifically, some residues mainly reappeared in 1H-15N HSQC spectrum, and some other residues particularly experienced an increase of peak height upon the addition of EquileninWe Enhancements in peak height by Equilenin was also calculated. The formula on the NMR signal enhancement (Ei) is almost similar with that of attenuation, but denominator and numerator are reversed. Fig. 4 shows Ei values as a function of amino acid residue number of KSI.). These features were also represented by? color on the three dimensional structure of KSI in Fig. 4 over a specific cut-off (levels divided by the averagewhich indicate that Equilenin was bound tightly to the active site of KSI, to reduce the effect of HyTEMPOon the protein.
4. Discussion
- HyTEMPO, a soluble and stable paramagnetic reagent, which has been known to cause a line broadening and not to convert the conformation ofproteins.[23,24] It offers the advantage of identifying the residues protected from the solvent. It is assumed from this experiment, that HyTEMPO may have entered into the hydrophobic cavity of KSI, to interact with nuclei of interest in KSI. Consequently, some of cross peaks in 1H-15N HSQC spectrum of KSI with HyTEMPO experienced attenuations in NMR signals. HyTEMPO especially leads to broaden the NMR signals of the hydrophobic residues in interior of protein remarkably affecting them. These characteristics of HyTEMPO can enable rapid mapping of binding site between the substrate and enzyme without three dimensional crystal structures.Equilenin an intermediate analog tightly binds to the active site of KSI by crystal packing interactions with neighboring molecules, although it was also exposed to the bulk solvent.[27] The dissociation constant (Kd) of the native KSI was determined to be about 2.6 M at pH 7.2 from a previous study.[28] Once Equilenin was accordingly added to KSI solution with HyTEMPO, NMR signals of many peaks dramatically were reappeared or strengthenedwhichmay attribute to the strong interaction between KSI and Equilenin. It means that most of HyTEMPO which is located in hydrophobic cavity dramatically appeared to be replaced by Equilenin from the hydrophobic cavity. The crystal structure of KSI with Equilenin was introduced and was analyzed as described before.[6] Residues such as Tyr16, Tyr32, Asp40, Tyr57, Trp92, Asp103, Trp120 in KSI play an important role about binding between KSI and Equilenin.[29] Furthermore, active site residues are surrounded closely by non polar groups (Met13, Ile17, Leu19, Val20, Ile25, Ile28, Val29, Met31, Ala36, Val38, Ile47, Ile53, Phe56, Leu61, Ala68, Met84, Phe86, Val101, Met105, Ile113, Met116, and Ala118).[6] As shown in Fig. 4, most residues in the middle and the end part of conical barrel of KSI were more experienced both by paramagnetic attenuations and enhancements than the upper part of those. Specifically, the residues which were disappeared in 1H-15N HSQC spectrum of KSI solution with HyTEMPO are spatially close to the active site residues of KSI. These observations can berelated to the size theory for HyTEMPO with Equileninwhich is approximately 4.4Å. Jang DS et al. reported that the distance between residues which formed Hydrogen bond was within 3Å.[25] Moreover, active site residues were tightly surrounded by substrate such as Equilenin. Hence, once Equilenin were injected to the KSI solution with HyTEMPO, polar active site residues in hydrophobic cavity of KSI are surrounded completely with non-polar of Equilenin. . As a result, the empty spaces distance for the hydrophobic cavity of KSI is within 4Å. The access of bulk solvent during enzyme catalysis is blocked by the layer of non polar residues and steroid substrate.[6] Consequently, once Equilenin is bound to KSI in solution with HyTEMPO, HyTEMPO comes out of hydrophobic cavity due to compactness of interaction between KSI and Equilenin.Thus, our NMR experiment using HyTEMPO as an indirect paramagnetic probe propose a potential method to find binding area of enzyme. It not only needs less time to obtain detailed information using NMR, but also need for structural information of protein is not required. Although this method can not describe the accurate active site between an enzyme and a substrate, but still it can provide rapid mapping for potential binding area between enzyme and a substratewhich can be used to study the functions of proteins irrespective to the use of three dimensional crystal structure for the same.
5. Conclusions
- In conclusion, we have investigated the potential active site for enzyme – substrate complex using KSI protein, HyTEMPO and multi-dimensional NMR technique. The peak height analysis revealed that HyTEMPO affects the hydrophobic cavity, particularly active site region. Some NMR signals experienced line broadening after injection of HyTEMPO. Meanwhile, many residues experienced the signal enhancements upon the addition of Equilenin on the KSI solution with HyTEMPO. Our result shows that potential active site region of protein can easily be detected by a paramagnetic reagent as an indirect probe using multi- dimensional NMR spectroscopy. Up till now, studies on active site mapping for proteins was mainly investigated by computational studies. This study is based on the experiment facts, and provides the best advantage of reduced time for the active site determination. Hence, this method enables the study of protein function without requirement of their three dimensional crystal structure.
 Abstract
Abstract Reference
Reference Full-Text PDF
Full-Text PDF Full-Text HTML
Full-Text HTML