-
Paper Information
- Paper Submission
-
Journal Information
- About This Journal
- Editorial Board
- Current Issue
- Archive
- Author Guidelines
- Contact Us
International Journal of Modern Botany
p-ISSN: 2166-5206 e-ISSN: 2166-5214
2014; 4(2): 48-60
doi:10.5923/j.ijmb.20140402.03
Role of Histone Deacetylases in Fungal Phytopathogenesis: A Review
Miroslava Cuperlovic-Culf1, Adrian S. Culf2
1National Research Council, Moncton, NB, Canada
2Atlantic Cancer Research Institute, Moncton, NB, Canada
Correspondence to: Miroslava Cuperlovic-Culf, National Research Council, Moncton, NB, Canada.
| Email: |  |
Copyright © 2014 Scientific & Academic Publishing. All Rights Reserved.
Acetylation of lysine is one of the major regulators of function of histone and non-histone proteins of eukaryotic cells, viewed as an avenue for cellular response to environmental, nutritional and behavioural factors. Aberrant protein acetylation as well as inhibition of acetylation or deacetylation has been related to variety of highly dissimilar diseases across species ranging all the way from cancers in humans to fungal diseases in plants. A number of small molecule secondary metabolites and non-ribosomal peptides from fungus as well as several secondary metabolites from plants have been suggested as deacetylation inhibitors. Inhibitory characteristics of a few of these compounds have been explored experimentally in detail and clinical uses have been investigated. For several others, their role in deacetylation inhibition has been suggested and shown in preliminary studies. Several of these inhibitory metabolites play a significant role in fungal phytopathology both as toxins and as metabolites engaged in protection. Present review establishes functions of Zn2+-dependent histone deacetylase enzymes of plants and fungi in phytopathology. The connection between metabolites and histone deacetylase enzymes and their reciprocal regulation particularly in relation to fungal invasion and plant resistance is also corroborated. The interaction between metabolites and representatives of different classes of histone deacetylases is reviewed that revealed binding characteristics of fungal cyclic peptides, fungal toxins, and plant secondary metabolites.
Keywords: Plant/fungus interaction, Acetylation/deacetylation, Histone deacetylases inhibitors
Cite this paper: Miroslava Cuperlovic-Culf, Adrian S. Culf, Role of Histone Deacetylases in Fungal Phytopathogenesis: A Review, International Journal of Modern Botany, Vol. 4 No. 2, 2014, pp. 48-60. doi: 10.5923/j.ijmb.20140402.03.
Article Outline
1. Introduction
- Emerging infectious diseases caused by fungi are recognized as a major threat to food production worldwide with observed growth in worldwide fungal infections in plants as well as other species [1]. Recently, the top ten fungal plant pathogens have been nominated [2]. These as well as a number of other fungal pathogens are leading to some of the most destructive diseases in plants including major damage to agricultural crops such as rice, wheat and corn. In addition to devastating plant yield, fungal toxins can present a danger to animal and human health through the consumption of infected plants with many historical examples of fungal toxins causing catastrophic losses of life [3]. The development of fungal infection resistant plants as well as novel anti-fungal methods is sorely needed. Phytopathogens such as fungi use a variety of life strategies. Fungi can directly enter plant epidermal cells, extend hyphae on top of, between or through plants cells or invaginate feeding structures into the plasma membranes of host cells. Plants lack mobile defender cells and a somatic immune system. Instead, they rely on an innate response of each cell and systemic signals released from infection sites [4]. Pathogen infection leads to activation of variety of responses in plant cells depending on the infection strategies and lifestyle of the pathogen as well as the type and resistance potential of the plant. Defence responses are associated with the expression of a wide range of genes, proteins and metabolites that can affect fungus directly, stimulate a cascade of processes in a specific cell or send signals across the plant to other cells. At the same time, fungi utilize a number of metabolites as toxins that are essential for their spread to the host. Regulation of the expression of these response/pathogenesis elements is in part performed by acetylation/deacetylation of proteins in plant or fungi. In turn, many of the toxins as well as resistance metabolites regulate the protein acetylation process. Acetylation regulation can cause changes in gene expression through regulation of histone proteins. Furthermore, it can directly activate or deactivate protein functions in different cell organelles through action on non-histone proteins. Acetylation has been known as a factor in transcription and protein activity regulation for 50 years [5]. Recently, an increasing number of publications have been showing a close link between metabolism and acetylation of proteins [6-9]. Protein acetylation has been shown to control toxin production in fungi as well as resistance response in plants [10]. Given the significance of the role of acetylation in plant/pathogen interaction, it is not surprising that several fungal toxins specifically target deacetylation regulator proteins – histone deacetylases (HDACs). Similarly, a number of plant metabolites target HDACs with important regulatory functions in biotic response. The evaluation and exploitation of natural metabolite regulation of resistance and pathogenesis is a very interesting approach for the development of phytopathogen resistant plants as well as bio- and natural product pesticides. In this review we will focus primarily on the interplay between metabolism and histone deacetylation in relation to plants’ biotic response to fungal infection as well as fungal pathogenesis. Regulation and interplay between metabolism and protein acetylation opens a wide range of new possibilities in agriculture as well as medicine.
2. Acetylation and Deacetylation in Pathological Fungus/Plant Interaction
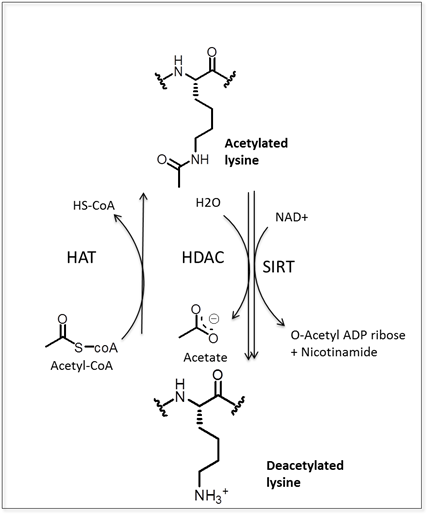
- Acetylation of lysine residues of proteins is a reversible modification regulated by the antagonistic activity of two groups of enzymes – histone acetyl transferases (HAT) and histone deacetylases (HDACs and SIRTs). HAT enzymes catalyze the transfer of acetyl groups from acetyl-CoA to the
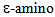 group of the lysine residue. HDACs and SIRTs remove the acetyl group from acetylated lysine residues releasing an acetate molecule in the presence of water (reviewed in Barneda-Zahoneroet al. [11] and Bannister and co-workers [12]) (Figure 1). Reversible acetylation is involved in regulation of highly diverse proteins including histones as well as a number of other nuclear, cytoplasmic and mitochondrial proteins [13, 14]. Chromatin modifications through acetylation and deacetylation of histone tails are directly involved in gene expression regulation. Positive charges on histone lysine residues are neutralized by acetylation leading to a relaxed conformation of chromatin. Relaxation of chromatin prevents generation of higher-order chromatin structures. These, more open chromatin chains, provide access for the transcription complex leading to gene expression [13]. Acetylated histones have been shown to be binding sites for bromodomain proteins which act as transcriptional activators [13]. Bromodomain specifically binds to acetylated lysine residues and is generally found in proteins that regulate chromatic structure and gene expression. Bromodomain proteins also bind to acetylated non-histone proteins that can act as transcriptional activators. Through gene expression regulation, acetylation of histones has been indicated as a regulator of many cellular processes including metabolism [14-18], fungal growth and pathogenesis [19] or plant hormone signaling [20].
group of the lysine residue. HDACs and SIRTs remove the acetyl group from acetylated lysine residues releasing an acetate molecule in the presence of water (reviewed in Barneda-Zahoneroet al. [11] and Bannister and co-workers [12]) (Figure 1). Reversible acetylation is involved in regulation of highly diverse proteins including histones as well as a number of other nuclear, cytoplasmic and mitochondrial proteins [13, 14]. Chromatin modifications through acetylation and deacetylation of histone tails are directly involved in gene expression regulation. Positive charges on histone lysine residues are neutralized by acetylation leading to a relaxed conformation of chromatin. Relaxation of chromatin prevents generation of higher-order chromatin structures. These, more open chromatin chains, provide access for the transcription complex leading to gene expression [13]. Acetylated histones have been shown to be binding sites for bromodomain proteins which act as transcriptional activators [13]. Bromodomain specifically binds to acetylated lysine residues and is generally found in proteins that regulate chromatic structure and gene expression. Bromodomain proteins also bind to acetylated non-histone proteins that can act as transcriptional activators. Through gene expression regulation, acetylation of histones has been indicated as a regulator of many cellular processes including metabolism [14-18], fungal growth and pathogenesis [19] or plant hormone signaling [20].
|
3. Metabolism Regulation by Protein Acetylation
- The majority of enzymes of intermediary metabolism are also regulated by acetylation [17, 23], thus making reversible lysine acetylation a route for direct metabolism control. Additionally, virtually all enzymes of central metabolism appear to be acetylated [18]. Acetylation and deacetylation processes are directly controlled by metabolism changes through substrates and co-factors for the acetylation process, e.g. acetyl-CoA and NAD+ making these two molecules a direct link between metabolism and protein acetylation. All HATs require acetyl-CoA as a donor of an acetyl group and are thus regulated by its concentration at the site of acetylation. SIRTs require NAD+ as a co-factor. Global reduction of nuclear acetyl-CoA levels decreases histone acetylation while reduction of NAD+ levels inhibits histone deacetylation by SIRT [7]. Acetyl-CoA, an acetate-thioester, is an activated form of acetate found in all organisms. Acetyl-CoA is produced in several different processes in cells, both in the cytoplasm and mitochondria [31]. In Saccharomyces cerevisiae and Candida albicans, acetyl-CoA is primarily produced from acetate by cytoplasmic acetyl-CoA synthetase, while in plants and animals acetyl-CoA can be derived from citrate [30]. In plant cells, acetyl-CoA is critical for the production of fatty acids, isoprenoids, flavonoids, phenolics, alkaloids and amino acid biosynthesis as well as the TCA and glyoxylate cycles and -oxidation. An increase of acetyl-CoA concentration stimulates genes that promote cell growth in yeast and these genes closely match those that are induced by c-Myconcoprotein in mammalian cells [6]. Acetyl-CoA levels influence acetylation with indications that a large, up to 10-fold, variation in the acetyl-CoA level in cells observed through the cell cycle profoundly and quickly affects acetylation levels [32]. Histone acetylation appears to be an extremely dynamic process with the half-life of histone acetylation possibly as short as 3 minutes [6, 33]. Histone acetylation in mammalian, as well as plant cells, also depends on the expression and function of ATP citrate lyase enzyme (ACL) [34]. Following growth factor stimulation and during differentiation, ACL converts glucose-derived citrate, obtained through partial Krebs cycle metabolism, into acetyl-CoA resulting in increased histone acetylation and gene expression. Therefore, glucose availability affects histone acetylation in an ACL-dependent manner. Increased histone acetylation promoted by amplified intracellular glucose levels leads to the expression of insulin-responsive glucose transporter (GLUT4), hexokinase-2 (HK2), phosphofructokinase-1 (PFK-1) and lactate dehydrogenase A (LDH-A), all significant regulators of glycolysis, leading to further increased glucose consumption and glycolysis as well as acetyl-CoA production [34]. As well as a factor in acetylation, acetyl-CoA is a major building block for the biosynthesis of fatty acids and sterols and is involved in isoprenoid-based protein modifications. In these roles, acetyl-CoA and ACL are a link between glucose and glutamine metabolism, fatty acid synthesis and mevalonate pathways as well as histone and protein acetylation regulation. Although histone acetylation is directly dependent on ACL, the acetylation of non-histone, cytoplasmic proteins, such as tubulin and p53, is not dependent on ACL function - at least in mammalian cells [35]. It appears that acetyl-CoA for non-histone protein acetylation can be obtained from a variety of pathways and sources rather than from just citrate in the ACL catalyzed pathway. However, further work is needed to establish these relationships in mammalian and even more non-mammalian cells.Global histone acetylation levels change across tissues and biological systems. They also change with metabolic or physiological alterations in the cells such as changes in pH levels [36]. A decrease in intracellular pH leads to global hypoacetylation of histones in an HDAC dependent manner [36]. As a result, there is an increased export of acetate anions and excess protons out of the cell leading to increased intracellular pH as well as decreased extracellular pH. This transport is performed by proton coupled monocarboxylate transporter (MCT). An increase in intracellular pH leads to the opposite effect with increased flow of acetate and protons into the cell and increased histone acetylation. In this process glucose, glutamine or pyruvate is required as a metabolic source for the production of acetyl-CoA. The relationship between intracellular and extracellular pH and acetylation levels for both histone and non-histone proteins deserve further attention, particularly in plant and fungal cells. Histone acetylation/deacetylation in fungi is crucial for the transcriptional regulation of a number of processes including secondary metabolism [37-41]. The first example of a direct link between histone acetylation and the expression of genes responsible for secondary metabolism in filamentous fungi was shown for the aflatoxin gene cluster in Aspergillusparasiticus [42]. Fungi treated with histone deacetylase inhibitors exhibit an increase in the production of diverse secondary metabolites suggesting that histone deacetylation acts as a regulator of biosynthetic pathways [43, 44]. Thus, further explorations of the interactions between histone acetylation and fungal secondary metabolism is needed for both control of toxic fungi and also for utilization of fungi for the production of known and novel chemical compounds.
4. Role of Plant HDACs in Resistance
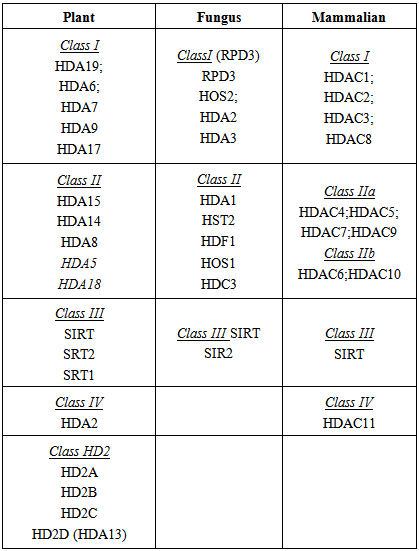
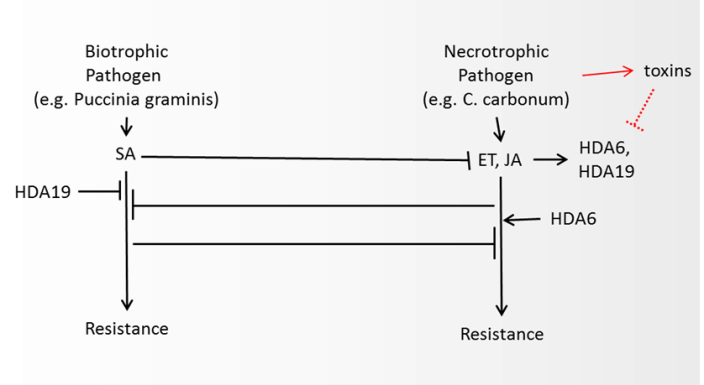
- Changes in the expression and activity of HDACs have been observed in relation to plant response to pathogens with distinct roles for different HDACs. Mutational and inhibition studies have indicated that HDACs control the expression and activation of a number of genes involved in the regulation of abiotic and biotic responses in different plants including regulation of the synthesis of plant hormones such as jasmonic acid, ethylene and salicylic acid. A schematic representation of our literature overview related to the known interactions between HDAC, salicylic and jasmonic acid pathways in relation to the response to biotrophic and necrotrophic pathogens (reviewed in Halim et al. [51] and Antico, et al. [52]) is shown in Figure 2. Biotrophic fungi require living plant tissue to survive. Necrotrophic fungi kill host tissues and feed on the remains. Synergistic or antagonistic interactions between these pathways, HDACs and their inhibitors allow the plant to optimize its response to a specific pathogen.
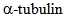 deacetylation. HDA14 similarly deacetylates
deacetylation. HDA14 similarly deacetylates 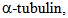 associates with
associates with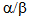 tubulin and is retained on GTP/ taxol-stabilized microtubules, in part through a direct association with the protein phosphatase 2A (PP2A). Similarly to HDAC6, HDA14 is found in both nucleus and cytoplasm. Association between HDA14 and PP2A suggests a link between protein phosphorylation and acetylation, both major regulators in biotic as well as other responses of cells [21]. Interestingly, inhibition of PP2A in a variety of plant species led to anti-fungal response, even in the absence of infection, suggesting that PP2A has a role in actively inhibiting the stress response in the absence of stress [63]. Due to PP2A’s association with HDA14 it can be hypothesized that HDA14 inhibition can influence the function of PP2A in a variety of processes including biotic stress response, although this hypothesis needs further experimental verification.Following literature-derived model schematically described in Figure 2, one can hypothesize that inhibition of HDA19 and HDA6 (Class I HDACs) leads to an enhanced salicylic acid (through the function PR proteins) and reduced jasmonic acid production. Inhibition of HDA14 possibly leads to an increased response of factors normally inhibited by PP2A. Therefore, one can hypothesize that necrotrophic pathogen progression benefits from HDAC inhibition, through its effect on the jasmonic acid pathway. On the other hand, resistance to biotrophic pathogens appears to be enhanced by HDAC inhibition. It should be kept in mind that complete HDAC inhibition as well as strong Class I HDAC inhibition leads to major changes in gene expression and can lead to cell death through autophagy or apoptosis as well as major changes in the function of ribozymes, mitochondria and other organelles.
tubulin and is retained on GTP/ taxol-stabilized microtubules, in part through a direct association with the protein phosphatase 2A (PP2A). Similarly to HDAC6, HDA14 is found in both nucleus and cytoplasm. Association between HDA14 and PP2A suggests a link between protein phosphorylation and acetylation, both major regulators in biotic as well as other responses of cells [21]. Interestingly, inhibition of PP2A in a variety of plant species led to anti-fungal response, even in the absence of infection, suggesting that PP2A has a role in actively inhibiting the stress response in the absence of stress [63]. Due to PP2A’s association with HDA14 it can be hypothesized that HDA14 inhibition can influence the function of PP2A in a variety of processes including biotic stress response, although this hypothesis needs further experimental verification.Following literature-derived model schematically described in Figure 2, one can hypothesize that inhibition of HDA19 and HDA6 (Class I HDACs) leads to an enhanced salicylic acid (through the function PR proteins) and reduced jasmonic acid production. Inhibition of HDA14 possibly leads to an increased response of factors normally inhibited by PP2A. Therefore, one can hypothesize that necrotrophic pathogen progression benefits from HDAC inhibition, through its effect on the jasmonic acid pathway. On the other hand, resistance to biotrophic pathogens appears to be enhanced by HDAC inhibition. It should be kept in mind that complete HDAC inhibition as well as strong Class I HDAC inhibition leads to major changes in gene expression and can lead to cell death through autophagy or apoptosis as well as major changes in the function of ribozymes, mitochondria and other organelles. 5. Role of Fungal HDACs in Pathogenesis
- Histone acetylation and other epigenetic processes are important regulators of fungal secondary metabolism [43, 44]. Histone modification of specific sites leads to activation of selected clusters of genes involved in a variety of metabolic pathways. This cascade can be initiated either by fungus itself or can be a target for reprogramming by other organisms. Treatment with the natural pan-HDAC inhibitor, trichostatin A resulted in increased production of a number of secondary metabolites in different fungi [43, 64]. Deletion or mutation of specific HDACs also led to changes in fungal growth and metabolism. HDF1 in Fusariumgramineaurum (class II) has been found to be an important factor for conidiation, sexual reproduction and pathogenesis in this pathogenic fungus that is responsible for fusarium head blight in wheat. Deletion of HDF1 also led to a significant reduction in virulence and toxin (deoxyvalenol, DON) production [19]. At the same time, deletion of the other two class II HDACs - HDF2 and HDF3- had no significant effect on conidiation and virulence showing that HDF1 is the major class II HDAC gene in the Fusariumgaminearum regulation of conidiation, DON production and plant infection. Earlier work by Ding et al. [65] has shown that deletion of the FTL1 gene in Fusariumgraminearum also results in reduced virulence while, at the same time, causing significant reduction in HDAC activity - once again suggesting an important role for HDAC in the virulence of this fungus.In Cochlioboluscarbonum (fungus producing HC Toxin), the gene HDC1, which is highly related to HOS2 in yeast (class I), is an important factor in virulence. HDC1 mutants had significantly reduced virulence due to reduced penetration efficiency, in spite of HC Toxin production at levels comparable to the wild type [66]. A similar effect was observed in Magnaportheoryzae, the most damaging fungal pathogen of rice. The infectious growth of this fungal species depends on the function of the complex Tig1, while at the same time deletion of any member of the Tig1 complex, which includes HOS2 (class I HDAC), lead to a significant reduction of HDAC activity. Once again, these results speak to the major role of HDAC activity in Magnaportheoryzae infection [65].The deletion of the class II HDAC, HdaA in A. nudulans and A. fumigatus promotes increased biosynthesis of penicillin and sterigmatocytin while, at the same time, affecting germination [39, 64]. In Fusariumfujikuroi, one of major pathogens of rice, FfHda1 (class II) and FfHda2 (class I) regulate secondary metabolism including the synthesis of toxins while FfHda4 (class II) is responsible for developmental processes. Deletion of both FfHda1 and FfHda2 makes Fusariumfujikuroi non-pathogenic [37].
6. Metabolic Inhibitors of HDAC Activity
- A number of pathogenic fungi make HDAC inhibitors either as their major pathogenesis-inducing metabolites or as metabolites of supporting virulence. Plants make molecules with HDAC inhibitory potential for either affecting fungal HDACs or for self-regulation as part of resistance to biotic or abiotic stressors. Reaction kinetics of acetylation and deacetylation depends on the concentration of the acetyl group in the proximity of the enzyme. At the same time, a number of primary and secondary metabolites can act as inhibitors of HDAC function. Lactate, the final product of glycolysis has been recently indicated as a potent inhibitor of HDAC activity [28]. Pyruvate, a product of glycolysis and a precursor for the TCA cycle, as well as a number of biosynthesis processes, is an ubiquitous HDAC inhibitor [25]. Apart from the major production of pyruvate in glycolysis, pyruvate is also synthesized during isochorismic acid synthesis [62] where it can be speculated that pyruvate plays a role in inhibiting HDA19 and releasing salicylic acid. Hait and co-workers [29] have shown one of the first examples of Class I HDAC regulation by metabolites. According to the result of Hait et al,. HDAC1 and HDAC2 are regulated by the endogenous lipid mediator sphingosine 1-phosphate (S1P). S1P is a bioactive sphingolipid metabolite formed by phosphorylation of sphingosine catalyzed by sphingosine kinase. S1P is an important signaling molecule found in plants, yeast, worms, flies and mammals [67, 68]. In nuclear extracts, S1P inhibited HDAC activity by 50% relative to 90% inhibition by a known, small molecule inhibitor (trichostatin A). HAT activity was not affected by S1P in these experiments. Interestingly, the structurally related molecules, sphingosine and lysophospholipid did not have any significant effect on HDAC1 or HDAC2. Also, out of all human HDACs, only HDAC1 and HDAC2 have been shown to bind to S1P. SphK2 – an enzyme that produces S1P in the nucleus from sphingosine is associated with HDAC1 and HDAC2 complexes providing proximal production of S1P to its nuclear target. Treatment of cells with an activator of protein kinase C that enhances activity of SphK2 led to nuclear export of SphK2 and transient inhibition of HDACs. Thus, S1P metabolism appears to be a signalling route for HDAC controlled gene expression and response to a variety of signals. S1P formed by nuclear SphK2 in response to environmental signals affected histone acetylation and gene expression through HDAC inhibition presenting a link between sphingolipidmetabolism in the nucleus and remodeling of chromatin and the epigenetic regulation of gene expression. In plant as well as yeast cells S1P has been also observed as an important signaling molecule possibly also serving a role in HDAC regulation [67]. In pathogenic fungi, lipid signaling has been indicated as an important factor in virulence as well as maintenance of cell wall integrity (reviewed in Singh and Del Poeta [69]). Sodium butyrate, a short chain fatty acid was the first natural product and metabolite indicated as an inhibitor of histone deacetylation [73] with a very interesting role in colon cancer prevention [74]. Also, sodium butyrate has been shown to inhibit the growth of human pathogenic yeasts: Candida albicans, Candida parapsilosis and Cryptococcus neoformans [75]. In plants, a derivative of butyrate,
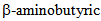 acid is an important metabolite previously observed in metabolomics analysis of wheat resistant to Fusarium sp. [76].
acid is an important metabolite previously observed in metabolomics analysis of wheat resistant to Fusarium sp. [76]. 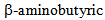 acid is in fact known as an important factor in the induction of resistance against pathogens in a variety of plants (recently reviewed by Justyna et al. [77]). The activity of
acid is in fact known as an important factor in the induction of resistance against pathogens in a variety of plants (recently reviewed by Justyna et al. [77]). The activity of 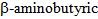 acid as an HDAC inhibitor has not been explored in any detail.
acid as an HDAC inhibitor has not been explored in any detail. 7. Specific Plant Metabolites Inhibiting HDAC
- Several plant metabolites have been indicated as possible HDAC inhibitors based primarily on their activity in human cells. Proposed HDAC inhibitors produced by a variety of plants include hydroxamic acids, short-chain fatty acids, boronic acids, α-keto acids, cyclic tetrapeptides, benzamides, ketones, isothiocyanates, organosulfur compounds, selenium-based compounds and their metabolites and other miscellaneous agents (reviewed by Rajendranet al. [70]). Some interesting examples include thujaplicin, diallyl disulfide, flavones, pomiferin and sulforaphane. Thujaplicin (isopropyl cycloheptatrienolones, Figure 3) is a natural monoterpenoid found in the wood of the trees from family Cupressaceae. Tropolone functional group in thujaplicin is known to be a strong metal ions chelate therefore making them possibly significant HDAC Zn2+ binders [109].
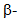 Thujaplicin has been shown as a very potent inhibitor of human HDAC2 and HDAC8 specifically [109]. It has shown anti-fungul activity against all examined wood-rotting fungi [108]. Further work is needed to explore its inhibitory potency against fungal and plant HDACs.
Thujaplicin has been shown as a very potent inhibitor of human HDAC2 and HDAC8 specifically [109]. It has shown anti-fungul activity against all examined wood-rotting fungi [108]. Further work is needed to explore its inhibitory potency against fungal and plant HDACs.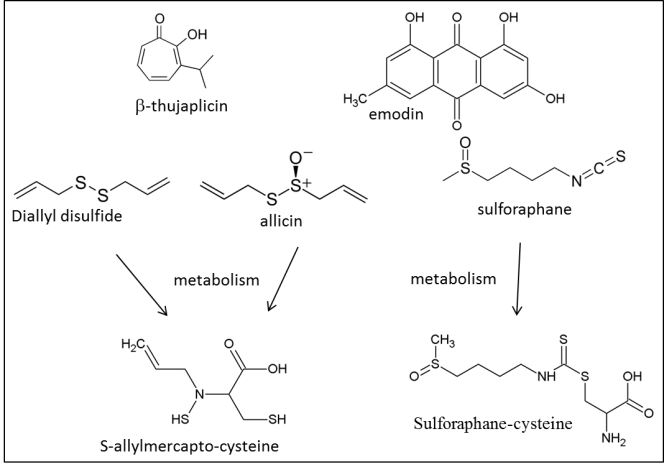 | Figure 3. Specific plant secondary metabolites with established HDAC inhibitory activity |
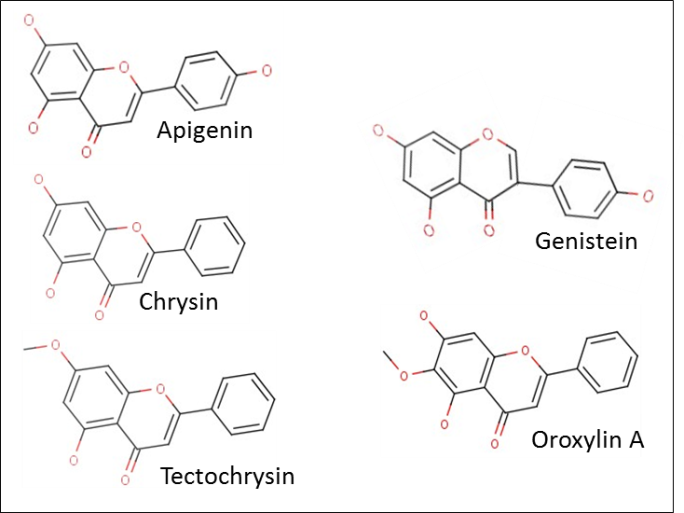 | Figure 4. Plant flavonoids with possible HDAC inhibitory activity suggested in literature |
8. Fungal Metabolites, Toxins and HDAC Inhibition
- Fungal secondary metabolites have specific host and protein targets that ensure the pathogen’s potency and specificity to a particular plant host. These metabolites, termed host-selective or host-specific toxins account for the specificity of host-pathogen interactions while making pathogens extremely effective. Several of these toxins, produced by necrotrophic fungi are now known to target HDACs. Fungal HDAC inhibitory toxins include trichostatin A [88], numerous cyclic tetrapeptides such as the apicidins [90], trapoxins [89], HC toxin and chlamydocin [91] and the linear molecule, depudecin [92]. The structures of these inhibitors are shown in Figure 5. TrichostatinA (TSA) was the first identified highly specific HDAC inhibitor [93] and its anti-fungal, anti-bacterial and anti-cancer properties have been proven many times since. Treatment of wheat seedlings with TSA has shown an overall rise in centromere association [94]. Recently, TSA testing in maize has shown the significance of HDAC regulation of cold resistance in plants [95]. Possible role of TSA in the suppression of fungal pathogens requires further exploration.Alternatia Brassicicola, a pathogen of the Brassica species, makes a small linear polyketide, HDAC inhibitor, depudecin [58, 96]. Depudecin contributes to the virulence of the fungus. However, its role appears to be much less important than the role of HC toxin on maize infection by C. Carbonum [96]. Depudecin non-producing mutants of pathogenic fungus Alternariabrassicicola have a small (10%) reduction in virulence on Brassica but not on Arabidopsis.Cyclic peptides are widely distributed secondary metabolites found in fungi as well as plants and lower animals [97] and they present the most important group of host-specific toxins. In fungi the cyclic peptides are synthesized using a template directed, nucleic-acid- independent nonribosomal mechanism [98, 99]. This synthesis is carried out by the multienzymenonribosomal peptide synthetase, the largest enzyme known in nature. Cyclic peptides produced by fungi have a variety of targets and roles with a subgroup showing very strong anti-HDAC activity. All currently known cyclic peptides from fungus with HDAC inhibitor roles have four amino acid residue macrocyclic rings and a tail, i.e. a linker with a zinc binding group. The structures of known examples of HDAC inhibitors of this form are shown in Figure 5.
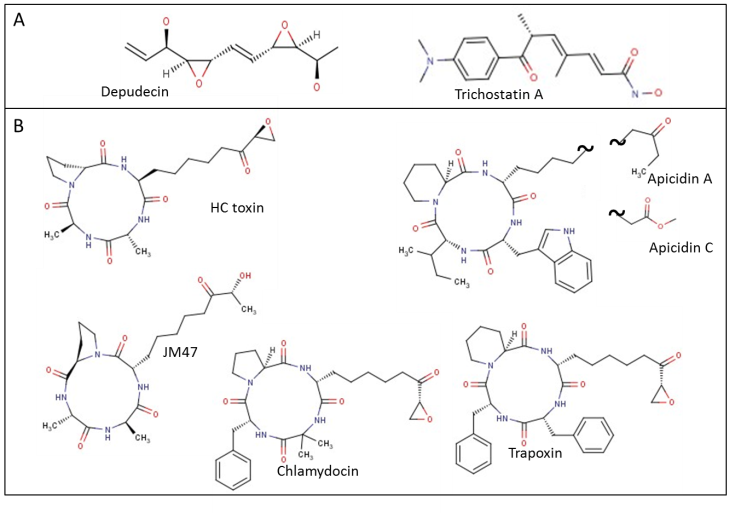 | Figure 5. Fungal secondary metabolites with known HDAC inhibitory effect. Shown are (A) small, linear molecules and (B) cyclic peptides |
9. Conclusions
- In both plant and pathogenic fungus, HDACs play an important role in their interaction. Several known necrotrophic fungi have highly potent HDAC inhibitors as major toxins which necessary for plant invasion. Plant HDAC inhibitors are generally much less specific and active - however in high concentrations they can have an effect on both plant and fungi HDACs. With HDACs having a significant role in plant development and biotic response as well as fungus pathogenesis further understanding the role of fungal and plant HDAC inhibitors as well as triggers for their production and their specific targets can lead to more effective, metabolite based fungus resistant plants or natural product fungicides. The increasing threat from fungi to human food sources should encourage research in this direction. Furthermore, in-depth analyses of the relationships between different classes of histone deacetylases in fungus and their secondary metabolism, growth and pathogenesis is essential for unlocking the mechanisms for synthesis of novel, natural product chemicals from fungi.
 Abstract
Abstract Reference
Reference Full-Text PDF
Full-Text PDF Full-text HTML
Full-text HTML