-
Paper Information
- Paper Submission
-
Journal Information
- About This Journal
- Editorial Board
- Current Issue
- Archive
- Author Guidelines
- Contact Us
International Journal of Genetic Engineering
p-ISSN: 2167-7239 e-ISSN: 2167-7220
2026; 14(1): 51-55
doi:10.5923/j.ijge.20261401.08
Received: Jan. 25, 2026; Accepted: Feb. 8, 2026; Published: Feb. 12, 2026

SSR-Based Evaluation of Fiber Quality and Color Traits in White and Naturally Colored Cotton F₄ Hybrids
R. B. Rakhmonova1, J. K. Umarova1, M. B. Khalikova2
1Navoi State University, Navoi, Uzbekistan
2Research Institute of Cotton Breeding, Seed Production and Agro-Technologies, Tashkent, Uzbekistan
Correspondence to: R. B. Rakhmonova, Navoi State University, Navoi, Uzbekistan.
| Email: |  |
Copyright © 2026 The Author(s). Published by Scientific & Academic Publishing.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/

This study was aimed at identifying the genetic factors controlling fiber quality and color in white- and naturally colored-fiber cotton genotypes and their derived F₄ hybrids. At present, the development of environmentally friendly cotton varieties with naturally colored fiber is considered one of the most important challenges, where the reduction in fiber quality and yield remains a major constraint. Therefore, the application of marker-assisted selection (MAS) technologies is of significant scientific and practical importance. In this study, genomic DNA was extracted from cotton genotypes and F₄ hybrids using the CTAB method. DNA quality and concentration were evaluated by agarose gel electrophoresis and a NanoDrop spectrophotometer. PCR analyses were performed using SSR markers BNL-1604, associated with fiber quality, and NAU-1043, associated with fiber color. The results demonstrated that both markers exhibited a high level of polymorphism. Allelic combinations detected with the BNL-1604 marker confirmed that several F₄ hybrids were promising in terms of fiber quality. The NAU-1043 marker revealed the inheritance pattern of fiber color and the presence of dominant alleles across generations. In conclusion, the findings indicate that the use of SSR markers enables early-stage evaluation of fiber quality and color in cotton breeding and facilitates the efficient selection of promising genotypes.
Keywords: Cotton, Fiber quality, Fiber color, SSR markers, F₄ hybrids, Marker-assisted selection (MAS)
Cite this paper: R. B. Rakhmonova, J. K. Umarova, M. B. Khalikova, SSR-Based Evaluation of Fiber Quality and Color Traits in White and Naturally Colored Cotton F₄ Hybrids, International Journal of Genetic Engineering, Vol. 14 No. 1, 2026, pp. 51-55. doi: 10.5923/j.ijge.20261401.08.
1. Introduction
- Cotton production is one of the leading sectors of agriculture, playing a crucial role not only in supplying raw materials for the textile industry but also in ensuring the economic stability of many countries. In recent years, global climate change, increasing environmental safety requirements, and the need for efficient use of production resources have necessitated the adoption of new scientific approaches in cotton cultivation systems [2,12]. From this perspective, the development of environmentally friendly cotton varieties requiring less energy during processing, particularly those with naturally colored fiber, has become one of the most pressing challenges [6].Naturally colored cotton varieties are distinguished by their ability to reduce the need for chemical dyes, minimize negative environmental impacts, and lower production costs [5]. However, numerous studies have shown that colored-fiber cotton genotypes generally lag behind white-fiber varieties in terms of fiber quality and yield performance [7,8]. In particular, the relatively low levels of key technological traits such as fiber length, strength, and uniformity hinder the large-scale industrial adoption of colored cotton [4].In conventional breeding programs, variety selection is primarily based on phenotypic traits. However, phenotypic expression is strongly influenced by environmental factors and may not fully reflect the true genetic potential of a genotype [1]. Moreover, economically important traits such as fiber quality and fiber color are typically polygenic in nature, making their inheritance difficult to assess solely through phenotypic evaluation [11].In recent years, the introduction of marker-assisted selection (MAS) technologies has significantly enhanced the efficiency of breeding programs [2]. The MAS approach enables the evaluation of genotypes at the DNA level and facilitates the early identification of genes or genomic regions associated with important agronomic traits [12]. Among the DNA markers used in this approach, simple sequence repeat (SSR) markers are particularly advantageous due to their high level of polymorphism, reliability, and cost-effectiveness [1,7].SSR markers have been widely applied to assess genetic diversity in cotton, determine genetic relationships among genotypes, and map quantitative trait loci (QTLs) associated with fiber quality and color traits [4]. Especially in colored cotton breeding, the identification of markers genetically linked to fiber color and quality plays a key role in accelerating the selection process [6].In the breeding process, the F₄ generation represents an important stage, as many heritable traits become relatively stabilized, increasing the possibility of identifying promising genotypes [11]. Nevertheless, hidden genetic variation may still persist in the F₄ generation and can only be detected using molecular marker [6]). Therefore, the analysis of F₄ hybrids based on SSR markers contributes significantly to improving breeding efficiency.In the present study, special attention was given to investigating the genetic factors controlling fiber quality and color in white- and naturally colored-fiber cotton genotypes and their derived F₄ hybrid combinations. The identification of genetic polymorphism using the fiber quality–associated BNL-1604 and fiber color–associated NAU-1043 SSR markers was aimed at selecting promising breeding materials. Previous studies have confirmed the genetic linkage of these markers with genomic regions controlling fiber quality and color traits [4,5].Unlike previous studies, this research focuses on early-generation (F₄) hybrids from Uzbek breeding germplasm, providing region-specific insights into fiber quality and color inheritance.Thus, this study is of high scientific and practical significance, as it addresses key challenges in colored cotton breeding, identifies genetic resources contributing to improved fiber quality and color, and supports the practical implementation of marker-assisted selection approaches in cotton improvement programs.
2. Materials and Methods
- Study object and plant materialThe study included 10 parental cotton genotypes representing white- and naturally colored-fiber groups selected from the national breeding program based on their contrasting fiber quality and color traits. White-fiber parents were characterized by superior fiber length (29–32 mm) and strength (28–32 g/tex), whereas colored-fiber parents possessed stable brown or green pigmentation but moderate fiber quality. Ten F₄ hybrid combinations derived from these parents were analyzed to evaluate segregation and stabilization of fiber quality and color traits.Fiber quality traits including fiber length (UHML), fiber strength, and micronaire were measured using HVI (High Volume Instrument). Fiber color intensity was visually scored and categorized into white, light brown, and dark brown classes. Phenotypic evaluation was conducted prior to molecular analysis to allow comparison between trait performance and SSR marker profiles.Genomic DNA extractionGenomic DNA was extracted from young leaf tissues using the classical CTAB method [3]. Leaf samples were ground in liquid nitrogen and incubated in CTAB extraction buffer. DNA was precipitated with isopropanol, washed with ethanol, and dissolved in sterile distilled water. This method is particularly effective for plants rich in polyphenols and polysaccharides, such as cotton, and allows the isolation of high-quality DNA [3].Assessment of DNA quality and concentrationThe quality of extracted DNA samples was assessed by 0.9% agarose gel electrophoresis, which allowed visual evaluation of DNA integrity and degradation [10]. DNA concentration and purity were determined using a NanoDrop™ 8 spectrophotometer by measuring A260/A280 absorbance ratios. Values ranging from 1.8 to 2.1 indicated high DNA purity (Thermo Fisher Scientific, 2018). For PCR analysis, all DNA samples were adjusted to a final concentration of 50 ng/µL.Selection of SSR markersSSR markers genetically associated with fiber quality and fiber color were selected based on published literature and information available in the CottonGen and NCBI (PubMed) databases [4,5]. As a result, the BNL-1604 marker associated with fiber quality and the NAU-1043 marker associated with fiber color were selected as the main diagnostic markers. These markers have previously demonstrated high levels of polymorphism and effectiveness in cotton breeding programs [8].
 | Table 1. Panel of DNA markers genetically associated with economically important traits in cotton (Gossypium hirsutum L.) |
3. Results
- The quality and concentration of genomic DNA extracted from parental forms and F₄ hybrid combinations were initially evaluated using agarose gel electrophoresis and a NanoDrop™ 8 spectrophotometer. Electrophoretic analysis showed that all samples were characterized by high-molecular-weight, non-degraded DNA bands, indicating that the extracted DNA was suitable for subsequent PCR analyses. Spectrophotometric measurements revealed A260/A280 ratios ranging from 1.96 to 2.08, confirming the high purity of the DNA samples. For PCR analysis, the concentration of all DNA samples was adjusted to 50 ng/µL.Clear and reproducible amplification products were obtained from all 20 cotton genotypes using the fiber quality–associated SSR marker BNL-1604. Electrophoretic analysis revealed three alleles with approximate sizes of 107 bp, 123 bp, and 135 bp, indicating a high level of polymorphism (Figure 1). The presence of multiple alleles within individual genotypes suggests that the BNL-1604 marker amplifies several homeologous loci.
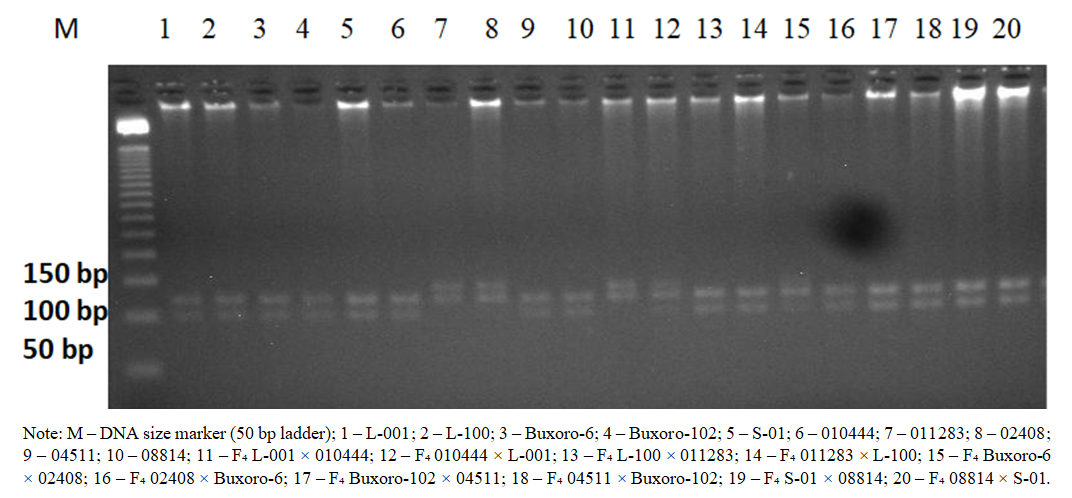 | Figure 1. PCR electrophoresis analysis of parental cotton genotypes and F₄ hybrids using the BNL-1604 SSR marker |
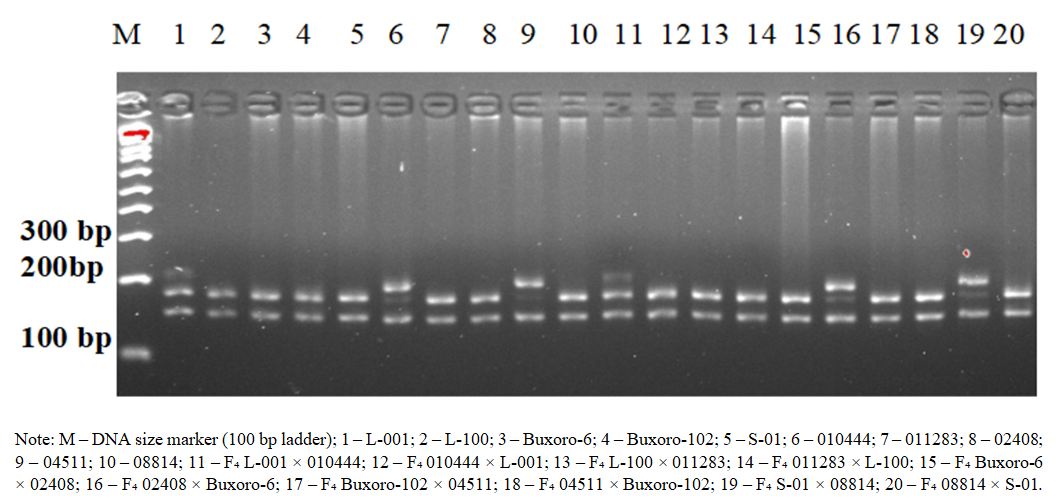 | Figure 2. PCR electrophoresis analysis of parental cotton genotypes and F₄ hybrids using the NAU-1043 SSR marker |
4. Discussion
- In this study, genetic factors controlling fiber quality and color were analyzed in white- and naturally colored-fiber cotton genotypes and their derived F₄ hybrid combinations using SSR markers. The obtained results once again confirm the effectiveness of molecular markers in cotton breeding and demonstrate the practical significance of the marker-assisted selection (MAS) approach.The high level of polymorphism detected for the BNL-1604 SSR marker indicates its strong genetic association with fiber quality. The identification of alleles with sizes of 107 bp, 123 bp, and 135 bp is consistent with previous studies reporting that the BNL-1604 marker amplifies multiple homeologous loci [4,7]. In particular, the presence of the 123 bp allele in all analyzed genotypes suggests that this allele is stable and widely distributed, making it a reliable diagnostic allele for fiber quality assessment.The simultaneous occurrence of the 107 bp and 135 bp alleles in several F₄ hybrid combinations indicates the preservation of genetic diversity at the molecular level. This finding confirms that although phenotypic stability may be achieved in the F₄ generation, a certain degree of genetic variation still persists. In this regard, the distinct performance of the F₄ hybrid combinations L-001 × 010444, 010444 × L-001, and Buxoro-6 × 02408 highlights their potential as promising genotypes for fiber quality improvement. Previous studies have also identified fiber quality–related QTL regions using BNL markers, and the present results further support these conclusions [8].The results obtained with the NAU-1043 SSR marker confirm the complex inheritance pattern of fiber color. The detection of alleles with sizes of 155 bp, 180 bp, and 200 bp, along with the simultaneous presence of three alleles in certain F₄ hybrids, indicates dominance effects and combined genetic influences on fiber color. These findings are consistent with earlier reports linking the NAU-1043 marker to fiber color–related QTL regions located on chromosome 7 [6].The dependence of fiber color inheritance on parental genotypes was also confirmed in the present study. The retention of dominant color-related alleles in specific hybrid combinations emphasizes the importance of selecting appropriate parental pairs in breeding programs. At the same time, the results demonstrate that it is feasible to improve fiber quality traits in colored cotton, indicating the possibility of simultaneous selection for both fiber color and quality.Overall, the findings show that the use of SSR markers serves as a powerful tool that complements—and in some cases surpasses—traditional phenotypic evaluation in cotton breeding. Through the MAS approach, early identification of promising genotypes associated with fiber quality and color can significantly shorten breeding cycles, optimize resource utilization, and facilitate the development of cotton varieties with high economic value [2,12].From this perspective, the results of the present study provide a solid scientific basis for informed decision-making in colored cotton breeding, the identification of genetically valuable breeding materials, and the future development of high-quality, environmentally friendly cotton varieties.
5. Conclusions
- The conducted molecular genetic studies enabled the effective identification of genetic factors controlling fiber quality and color in white- and naturally colored-fiber cotton genotypes and their derived F₄ hybrid combinations using SSR markers. The high quality of the extracted genomic DNA ensured the reliability of PCR analyses.The high level of polymorphism detected with the fiber quality–associated SSR marker BNL-1604 demonstrated its significant diagnostic value in cotton breeding. The simultaneous occurrence of the 107 bp, 123 bp, and 135 bp alleles in several F₄ hybrids confirmed the presence of genetically superior combinations with respect to fiber quality. In particular, the F₄ hybrids L-001 × 010444, 010444 × L-001, and Buxoro-6 × 02408 were identified as promising breeding materials.The results obtained using the NAU-1043 SSR marker revealed that fiber color exhibits complex inheritance patterns that are strongly dependent on parental genotypes, emphasizing the importance of selecting appropriate parental combinations in colored cotton breeding programs. Overall, the findings demonstrate that marker-assisted selection (MAS) based on SSR markers is an effective approach for the early identification of promising genotypes in cotton breeding.
 Abstract
Abstract Reference
Reference Full-Text PDF
Full-Text PDF Full-text HTML
Full-text HTML