-
Paper Information
- Paper Submission
-
Journal Information
- About This Journal
- Editorial Board
- Current Issue
- Archive
- Author Guidelines
- Contact Us
International Journal of Agriculture and Forestry
p-ISSN: 2165-882X e-ISSN: 2165-8846
2017; 7(3): 57-66
doi:10.5923/j.ijaf.20170703.01

Genetic Structuring of Four Tectona grandis L. f. Seed Production Areas in Southern India
Sreekanth P. M.1, Balasundaran M.2
1Department of Biotechnology, Cochin University of Science and Technology, Kochi, Kerala, India
2Environmental Biotechnology, Tropical Institute of Ecological Sciences Ecological Research Campus, Kottayam, Kerala, India
Correspondence to: Sreekanth P. M., Department of Biotechnology, Cochin University of Science and Technology, Kochi, Kerala, India.
| Email: |  |
Copyright © 2017 Scientific & Academic Publishing. All Rights Reserved.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/

Teak (TectonagrandisL. f.) is a tree species indigenous to India and other South and East Asian countries. It produces high-value timber and is easily established in plantations. Reforestation requires a constant supply of high quality seeds. Seed production areas (SPA) of teak are improved stands used for collection of open-pollinated quality seeds in large quantities. Information on the genetic diversity of major teak SPAs in India is scanty. In this study we analyzed the genetic structure of four important seed production areas of Kerala State in south India by employing amplified fragment length polymorphism markers using ten selective primer combinations on 80 samples (4 populations x 20 trees). The gene diversity of the SPAs varied from 0.18 (Konni SPA) to 0.22 (Wayanad SPA). The percentage of polymorphic loci ranged from 61.3 (Parambikulam SPA) to 72.9 (Nilambur SPA). The mean combined gene diversity index (HT) of all four SPAs was (HT = 0.2395; SE 0.0100; p < 0.0001). Analysis of Molecular Variance (AMOVA) based on the two genetic clusters indicated that majority of genetic variation (75%) occurred within populations, while the variation between the two clusters was 25%. Nei’s genetic differentiation (GST = 0.17) Bayesian estimates (FST = 0.1433 SE 0.0436; P < 0.0001) supported the AMOVA partitioning and provided greater genetic diversity within seed production areas. A UPGMA dendrogram was prepared and showed that all twenty samples from each of Konni and Parambikulam SPAs clustered into two separate groups, respectively. However, five Nilambur genotypes and one Wayanad genotype intruded into the Konni cluster. The higher gene flow estimated (Nm = 2.4) reflected the inclusion of Konni origin planting stock in the Nilambur and Wayanad plantations. Evidence for population structure investigated using 3D Principal Coordinate Analysis of FAMD software 1.30 indicated that the pattern of clustering was in accordance with the origin of SPAs. The AFLP fingerprinting was capable of identifying the geographical origin of planting stock and thereby revealing the occurrence of errors in genotype labeling.
Keywords: AFLP, Teak, Genetic structure, Seed production area
Cite this paper: Sreekanth P. M., Balasundaran M., Genetic Structuring of Four Tectona grandis L. f. Seed Production Areas in Southern India, International Journal of Agriculture and Forestry, Vol. 7 No. 3, 2017, pp. 57-66. doi: 10.5923/j.ijaf.20170703.01.
1. Introduction
- Teak (Tectona grandis L. f.) is indigenous to India and other South and East Asian countries. This species produces high-value timber, is fast growing and is easily established in plantations. These factors make teak one of the most promising species for sustainable production in the tropics [1] [2]. Teak is widely planted in commercial plantations in areas of its original range of tropical Asia and as an introduced species in Africa, Central and Latin America. Teak plantations account for 5 to 8 percent of the total forest area in the tropics [3] and comprise 90 percent of the quality hardwood plantations raised for timber production [4]. Presently, about 1.5 million ha of teak plantations exist in India and around 50,000 ha are replanted annually [5]. After completion of one rotation period, teak plantations are clear-felled and then replanted with seedlings raised from improved seed sources, such as seed orchards and seed production areas (SPAs). A seed orchard is a plantation of selected clones or progenies protected from pollen contamination from outside the orchard, and managed to produce abundant and easily harvested seed crops [6]. Generally, seed production areas are stands of suitable provenances raised from seeds of a reasonably large number of unrelated trees and managed specifically for seed production. But the quickest and the most common method adopted to establish a seed production area is to convert an existing stand of good growth and genetic quality [7]. Such SPAs are highly relevant for long-rotation crops like teak since they assure immediate availability of adequate quantities of seeds. In India, the first teak plantation was planted along the banks of river Chaliyar in Nilambur, Kerala, south India, during 1842 (Fig. 1). About 2.28 ha of this original plantation still exist as a permanent preservation plot. At present, 75,300 ha of teak plantations are spread over various forest divisions of Kerala, including Wayanad, Nilambur, Parambikulam, Achenkovil, Konni, Ranni and Malayattoor [8]. There is no record of origin of planting stock, but it is likely that earliest teak plantations originated from seeds collected from nearby natural forests. The harvestable age of teak ranges from 50 to 60 years in different forest divisions. An average of 1000 ha of teak plantations are clear felled and replanted each year after reaching harvestable age in Kerala. Since seed availability from clonal seed orchards is extremely poor, the nursery stock (stump/root trainer seedlings) is raised using seeds collected from plantations converted to SPAs [9]. An SPA of teak is usually referred to as a phenotypically superior stand comprising vigorously growing trees, upgraded by thinning and culling infected and poorer phenotypes, and then managed to produce abundant seeds. So far 5,000 ha of SPAs have been established in India [10], of which, about 1250 ha are in Kerala [11].
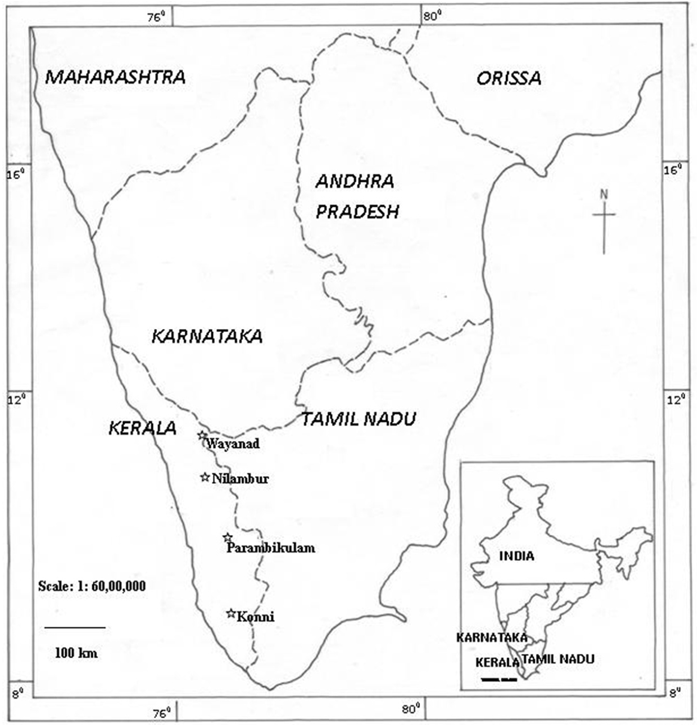 | Figure 1. Map showing the location of the four seed production areas of Kerala selected for AFLP analysis |
2. Materials and Methods
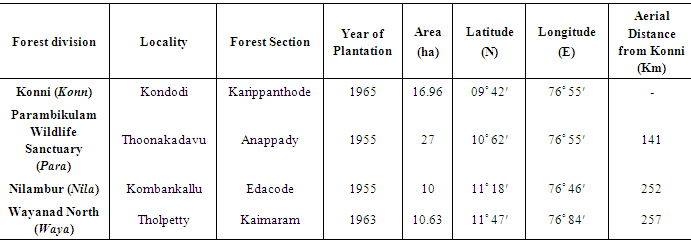
- Four younger SPAs (planted after 1954) located in four important geographic areas of teak plantations namely, Konni (Konn), Parambikulam (Para), Wayanad (Waya) and Nilambur (Nila) were selected for study (Fig. 1, Table 1). From each SPA, 20 trees were selected by using a line transect method. Sample trees were selected at intervals to collect 20 samples along the transect that was drawn at the broadest area of the plantation so as to obtain a true representation of the entire SPA. From these 20 trees, expanding leaves were collected for DNA extraction.
|
3. Results
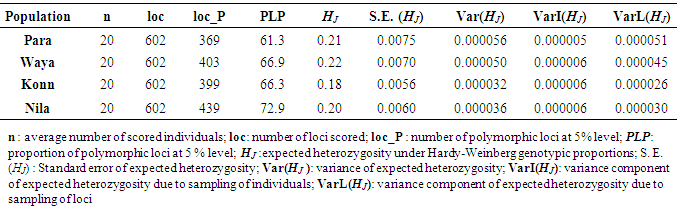
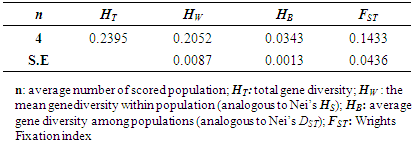
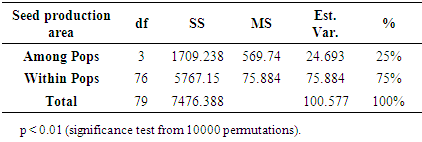
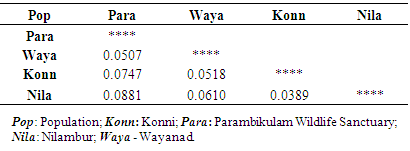
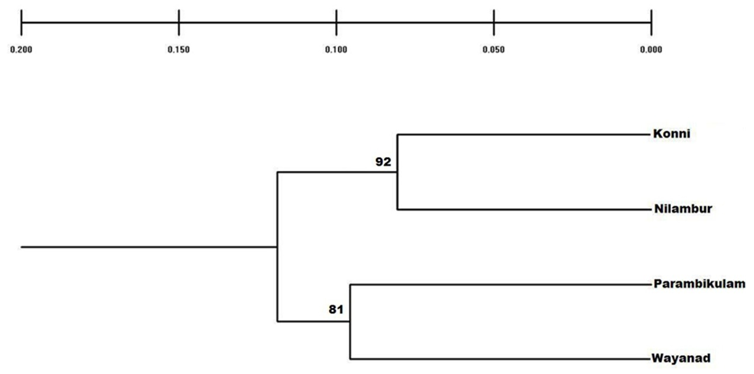
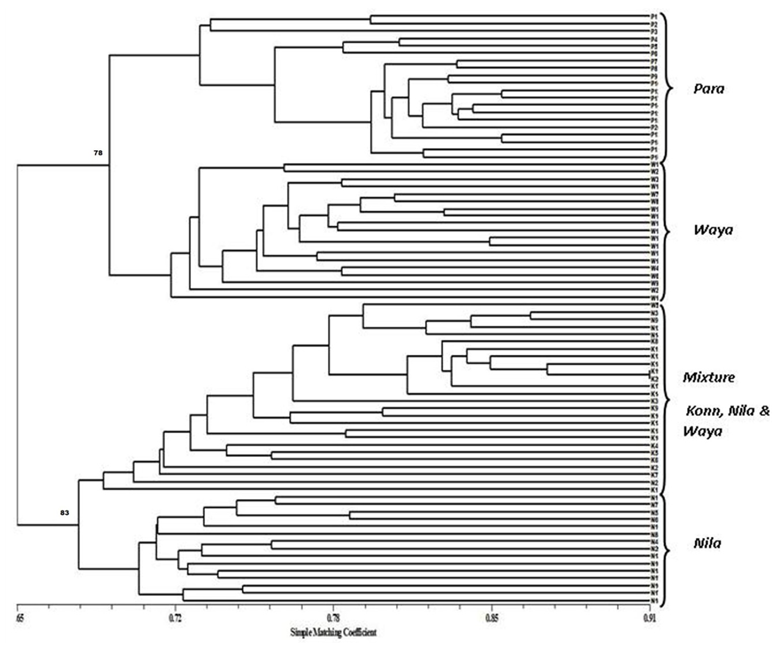
- Genetic diversity within and among populationsSix hundred and two loci were scored for statistical analysis. The percent of polymorphic loci (PPL) ranged from 61.3 (Parambikulam) to 72.9 (Nilambur). Nei’s gene diversity (H) or expected heterozygosity (HJ) ranged from 0.18 (Konni) to 0.22 (Wayanad) (Table 2). The average gene diversity within populations (HS = 0.2052; SE 0.0087; p < 0.0001), the total gene diversity (HT = 0.2395; SE 0.0100; p < 0.0001) and Wrights Fixation index (FST = 0.1433 SE 0.0436; P < 0.0001) were calculated for overall set of populations. The relative magnitude of Nei’s genetic differentiation among subpopulations, GST, was 0.17 indicating that 17 percent of total gene diversity was recorded among the four SPAs while the remaining 83% was recorded within the SPAs. The indirect measure of gene flow estimated from the GST value was 2.4. Results from three analyses (Bayesian FST = 0.1433; Nei GST = 0.17 and AMOVA (Table 4) showed greater variability within the seed production area.UPGMA dendrogram and PCoAUPGMA clustering showed that Konni and Nilambur SPAs are separated with least genetic distance (0.0389), Parambikulam and Nilambur SPAs showed the greatest genetic distance (0.0881) (Table 5). An UPGMA dendrogram was constructed to group the four SPAs based on genetic distance. The dendrogram separated Nilambur and Konni SPAs from Parambikulam and Wayanad SPAs. Bootstrap support values are provided at the corresponding node for each cluster (Figure 2).The cluster analysis comprising 80 trees revealed a unique dendrogram. The dendrogram divided 80 genotypes into two main clusters. The first cluster was shared between genotypes from Parambikulam to Wayanad; the genotypes of the SPAs formed their own sub clusters based on their geographic location. The second cluster was shared between Nilambur and Konni SPA genotypes but with mixing of genotypes from two geographic areas in one sub-cluster. One of the clusters consisted of 15 genotypes, all from Nilambur SPA, while the second cluster comprised of five genotypes from Nilambur (N2, N3, N9, N12, and N14) and one from Wayanad (W5) in addition to all 20 genotypes from Konni. This showed that these five genotypes from Nilambur and one from Wayanad might have actually originated from Konni. Bootstrap support values are provided at the corresponding node for each cluster (Figure 3). The PCoA revealed the genetic relationships between populations used in this study (Figure 4). The scale plot obtained for PCoA analysis showed similar genetic structuring of populations based on the origin of genotypes of the SPAs.
|
|
|
|
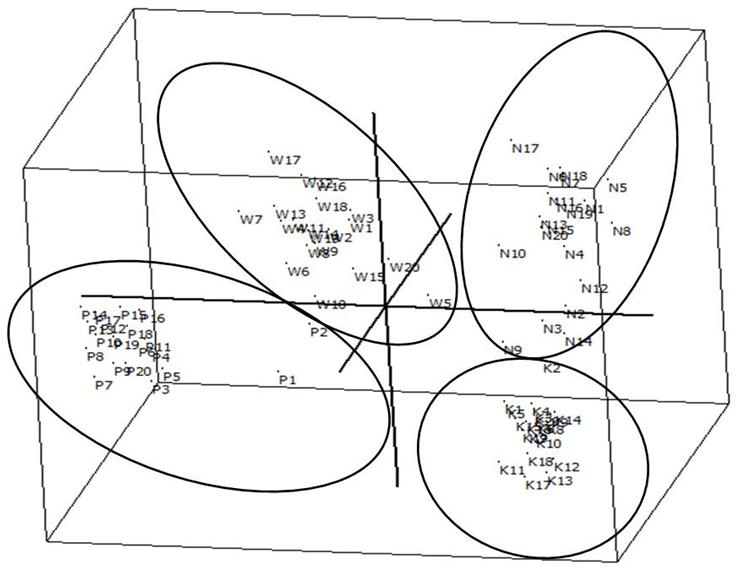 | Figure 4. 3D principal co-ordinate analysis (PCoA) of 80 genotypes of the four seed production areas performed using FAMD 1.30 software |
4. Discussion
- Ten AFLP primer combinations showed a total of 602 loci. The dominant markers of 100-200 number provides consistent estimate to evaluate the genetic diversity parameters [34]. Sample sizes above ten individuals are required to estimate marker-based within-population diversity [35, 36]. AFLP markers 238.1 ± 227.9 employed to an average of 14.0 ± 14.8 populations and 14.5 ± 9.0 individuals per population efficiently perform within and between population genetic diversity analysis in plants [37]. The present investigation was based on four important seed production areas each with 20 individuals i.e. 20 ± 0.0 individuals per location (Table 1) used to analyze the preliminary genetic diversity data within and between the teak seed production areas of Kerala.A similar study conducted using AFLP markers on teak trees growing in natural forests of the same geographic regions reported the genetic diversity of the four provenances from which the SPAs might have originated [38]. The genetic diversity indices of all four natural forest teak populations were higher than those for the SPAs that were established in the respective geographic region of the Western Ghats. The total gene diversity index of the four SPAs was 0.2395 ±0.0201, lower than the total gene diversity of the nine natural teak populations (0.2644 ±0.0178) reported in the previous study which covered geographic regions from two additional states of Karnataka and Tamil Nadu. Since teak is a cross pollinated species, reduction in genetic diversity in a randomly mating population could exert adverse impacts not only on seed production but also on crop productivity. Surendran et al. [39] considered selective culling of inferior phenotypes from a seed stand as a ‘negative selection’ sufficient for 5-10% genetic gain. However, studies have not been undertaken of the potential or realized genetic gains from negative selection in teak SPAs or the impacts on seed production caused by reduction in genetic diversity of SPAs. The overall gene diversity index value for Kerala SPAs is lower than the values from other wind pollinated, woody long lived tree species [40]. Examples are Populus tremuloides (0.30), P. grandidentata (0.35) and Quercus petrea (0.298) [41]. Using RAPD analysis Prabhu [12] reported that genetic diversity was low in teak SPAs of Kerala located beyond my study area. The reason for the low diversity in teak SPAs could not be identified. The plantations might have been raised using seeds of a couple of mother trees of narrow genetic base. Such progenies are likely to show inbreeding depression and poorer performance than their parents [7]. Genetic differentiation and gene flow among populationNei’s genetic differentiation (GST = 0.17), Wrights fixation index (FST = 0.143) (Table 3), and AMOVA (Table 4) all showed greater genetic diversity within SPAs than between SPAs. Genetic diversity status of plus trees of teak have been studied using biochemical (isozymes) and molecular (RAPD/AFLP) markers in recent years (Kertadikara and Prat, [42]; Kjaer and Suangtho, [43]; Changtragoon and Szmidt, [44]; Nicodemus et al. [45]; Shrestha et al. [46]; Sreekanth et al. [38]. These studies confirmed that teak is an out-breeding species and revealed maximum genetic variation within as opposed to between populations. Higher levels of population differentiation (between-population diversity) have also been reported in other tropical tree species screened using RAPD markers as in Gliricidia sepium (40%) (Chalmers et al. [47] and in Swietenia macrophylla (20%) (Gillies et al. [48], both of which are entomophilous and extreme outbreeders like teak. Generally long-lived and outcrossing species tend to be more genetically diverse and have less genetic differentiation between their populations [49].Clustering and PCoA of individual genotypes from SPAThe pattern of clustering of the individual genotypes was in accordance with the geographic origin of SPAs as determined in the previous study on natural teak populations [38]. All 20 samples from Konni and from Parambikulam SPAs clustered into respective groups clearly justifying their respective geographic origin. However, five genotypes namely, N2, N3, N9, N12 and N14 from Nilambur and one Wayanad genotype, W5, intruded into the Konni cluster. I could not able trace the origin of the seed as there was no information available from the forest offices. Records such as Plantation Journal were also unavailable for reference. The depths of genealogy within SPAs depend on several population genetic parameters, including effective population size, natural selection and recombination. Genes such as immunity genes (Klein et al. [50], and self-incompatibility [51] are highly prone to lineage sorting due to over dominant selection. In such cases, it is much less likely to maintain shared haplotypes between species based on geographical location [52]. The indirect estimate of gene flow obtained from the AFLP marker study was Nm = 2.4. This is substantial when compared to teak in the natural forests (Nm = 1.5) of the Western Ghats [38]. By definition gene flow is the movement of genes from one generation to another or from one place to another through pollen movement or through seeds. Geographic variation of natural teak populations has been detected in Kerala [38]. Seed transfer guidelines should be developed on the basis of genetic information and data from provenance trials. The level of genetic diversity is also crucial to ensure the long-term success of a restoration project [53]. If commercial plantations are derived from a few superior trees / plus trees, their genetic diversity tends to be low. If the gene flow from seed production area into native plantations is severe, the distinct gene pools of native forests will be destroyed [54]. Seeds used for plantations may be collected from a few seed trees that reveal capable phenotypes and are grown in nurseries [55, 56], avoiding the selection pressure on seed establishment. In the UPGMA clustering and PCoA analysis Konni clustered with Nilambur by virtue of their genetic similarity even though these locations are geographically far apart. The seed production areas are often established in close proximity to native inter-fertile conspecific populations using seeds from non-local and sometimes distant sources. One of the possible negative concerns of this planting policy is artificial genetic homogenization, the increase in the genetic similarity between the introduced and native gene pools with time via gene flow [57]. Especially in native populations of relatively small size, the human-mediated inflow of non-local genes may eventually swamp locally adapted genotypes. This process has rarely been assessed in natural ecosystems. This study showed the utility of AFLP fingerprinting in identifying the origin of planting stock and thereby revealed the occurrence of the mistake of genotype identity. Similar mislabeling errors were detected by using AFLP markers to study a clonal seed orchard of teak established in 1985 at Kulathupuzha in southern Kerala [58]. These seed production areas are major provenances in Kerala for the quality and quantity of seeds and often established in close proximity to reserve forests. Since there is no record on the origin of seeds during the establishment of these seed production areas the genetic composition of these commercial plantation was genetically different from that of natural populations, gene flow from commercial plantations into native forests would pose serious problems.
ACKNOWLEDGEMENTS
- The authors acknowledge the financial support provided by the Department of Biotechnology, Government of India (S.O.No.BT/PR/2302/AGR/08/161/2000). We are thankful to the Forest Departments of Kerala for giving permission for the collection of leaf materials.
 Abstract
Abstract Reference
Reference Full-Text PDF
Full-Text PDF Full-text HTML
Full-text HTML