-
Paper Information
- Paper Submission
-
Journal Information
- About This Journal
- Editorial Board
- Current Issue
- Archive
- Author Guidelines
- Contact Us
American Journal of Bioinformatics Research
p-ISSN: 2167-6992 e-ISSN: 2167-6976
2014; 4(2): 23-32
doi:10.5923/j.bioinformatics.20140402.01
A Comprehensive in silico DNA Sequence Analysis of Sperm Surface ADAM Genes Collected from RefSeq Database
Ayman Sabry 1, 2, Alaa Ahmed Mohamed 1, 2, 3, Nabil Said Awad 1
1Biotechnology and Genetic Engineering Unit, Scientific Research Deanship, Taif University, TAIF, Zip Code 21974, Post Box 888 KSA
2Cell Biology Department, National Research Cen- ter, Dokki, Giza, Egypt
3Animal Reproduction and AI, Veterinary Research Division, National Research Center, Dokki, Giza, Egypt
Correspondence to: Ayman Sabry , Biotechnology and Genetic Engineering Unit, Scientific Research Deanship, Taif University, TAIF, Zip Code 21974, Post Box 888 KSA.
| Email: |  |
Copyright © 2014 Scientific & Academic Publishing. All Rights Reserved.
The objective of the present study is to provide a comprehensive in silco sequence analyses of sperm-surface ADAM genes using the curated and nonredundant RefSeq database. 36 complete refseq CDS (coding sequence) of 9 members of ADAM gene family namely ADAM1,2,3,4,5,6,18,24 and 32 were obtained to investigate its evolution and differentiation within and among species. Among the 9 sperm-surface genes ADAM1 has the longest CDS length, the highest GC% and largest variation in base composition. Measurement of polymorphism and genetic diversity (e.g. Number of haplotypes, nucleotide diversity (π) and average number of nucleotide diversity) varied greatly among the sperm-surface ADAM genes. These large variations are evidence for the effect of selection pressure on these genes. The phylogenetic analysis displayed clearly the resolved relationship of sperm-surface ADAM genes and most of bootstrap support were high. Sperm-surface ADAM genes were classified into 6 clades where, ADAM1,2,3,4,5,6,18,24 and ADAM32 of B.taurus and H. sapiens categorizing to one large lineage where ADAM32 of R. norvegicus and M. musculus belonging to another separate lineage. In conclusion, the in silico analysis of the 9 sperm-surface ADAM genes showed a great deal of variation among and within this genes indicating the presence of localized signals of selection pressure on these genes. Moreover, ADAM1,4,6 and 24 have no human orthologues. More importantly, our results suggested that ADAM1,2,3,4,5,6, 18,24 and ADAM32 of B.taurus and H. sapiens were descendent form one ancient ancestor, where ADAM32 of R. norvegicus and M. musculus have another ancestor.
Keywords: Sperm-surface, ADAM genes, in silco Sequence analysis, DNA sequence diversity, RefSeq
Cite this paper: Ayman Sabry , Alaa Ahmed Mohamed , Nabil Said Awad , A Comprehensive in silico DNA Sequence Analysis of Sperm Surface ADAM Genes Collected from RefSeq Database, American Journal of Bioinformatics Research, Vol. 4 No. 2, 2014, pp. 23-32. doi: 10.5923/j.bioinformatics.20140402.01.
1. Introduction
- The interaction of mammalian spermatozoon with the oocytee’s extracellular matrix or zona pellucid is critical first step towards successful fertilization. Important key players in this extracellular interaction are ADAM (A Disintegrin And Metalloproteinase) genes. The ADAM gene family comprises 35 characterized genes in mammals, with about 18 genes are known to be expressed exclusively or predominantly in the male reproductive tissue (Wolfsberg et al., 1995b, Blobel, 1997, Black & White, 1998, Primakoff & Myles, 2000, Seals & Courtneidge, 2003, Cho, 2005 and Grayson & Civetta, 2013). ADAM gene family was unearthed during the study of sperm and egg merger that start off zygote development (Blobel et al., 1992, Wolfsberg et al., 1993 and Wolfsberg et al., 1995a). ADAMs were found to have multiple and diverse functions, both tissue-specific as well as ubiquitous patterns of expression and common evolutionary history (Finn & Civetta, 2010). The analysis of ADAM family evolution among mammals has found faster divergence of genes expressed in testes (Civetta, 2003, Glassey & Civetta, 2004, Finn & Civetta, 2010 and Morgan et al., 2010).In mammals ADAM1,2,3,4,5,6,18,24 and 32 are best characterized in terms of their role during fertilization. The first three genes play a role during sperm migration and zona pellucida (ZP) binding as well as egg membrane recognition and fusion (Blobel et al., 1992, Wolfsberg et al., 1995a, Primakoff & Myles, 2000 and Evans, 2002). For example, knockout mice for ADAM2 and ADAM3 show drastic decreases in sperm aggregation, a trait that has been suggested to confer sperm with competitive advantages (Moore et al., 2002, Fisher & Hoekstra, 2010 and Han et al., 2010). ADAM3 knockouts males were infertile due to deficiencies in sperm-ZP interactions, and more importantly, sperm migration into the oviduct (Shamsadin et al., 1999, Nishimura et al., 2001 and Yamaguchi et al., 2009). ADAM2 knock- outs also significantly affect reproductive success. In vivo, ADAM2 null mice have a fertility rate 50 times lower than the wild-type. This drop in fertility once again does not appear to be the result of a single process, but is instead a combination of deficiencies in sperm-egg fusion, sperm-egg binding, spermZP binding and sperm migration (Cho et al., 1998). ADAM1a knockouts result in sperm unable to migrate to the egg; in vivo the knockout produces an infertile phenotype but in vitro, sperm are able to fertilize eggs. ADAM1b knockouts appear to produce normal sperm but affect the levels of ADAM2 on mature sperm (Nishimura et al., 2004 and Kim et al., 2006).ADAM4,5,6,18,24, and 32 genes are also sperm surface genes while other ADAMs have been identified only as testis expressed (Frayne et al., 2002, Kim et al., 2005 and Zhu et al., 2009). Six of the sperm surface genes (Adams 1 to 6) assemble into functional complexes. Currently, there is evidence for three sperm-specific complexes (ADAM2- ADAM3- ADAM4, ADAM2-ADAM3-ADAM5, and ADAM2-ADAM3- ADAM6), two testis-specific complexes (ADAM1a- ADAM2, and ADAM2-ADAM3), and one complex common to both (ADAM1b-ADAM2) (Cho, 2012). All complexes require at least ADAM2 and/or ADAM3, if not both, and their interactions appear to be central for a variety of sperm functional adaptations to fertility in mice.From molecular evolutionary stand point, most of protein coding genes have been found to evolve under purifying selection, but genes that function in perception, immunity and reproduction are often fast-evolving exceptions to this rule (Voight et al., 2006, Kosiol et al., 2008 Koonin & Wolf, 2010. Reproductive genes, such as those that code for species-specific fertilization proteins, male accessory gland proteins, and sperm proteins have been shown to exhibit rapid evolution in taxa as diverse as invertebrates, mammals and plants (Swanson & Vacquier, 2002, A, 2003, Clark et al. , 2006, Panhuis et al., 2006, Turner & Hoekstra, 2008 and Dorus et al., 2010).One of the distinguished NCBI projects is Reference Sequence (RefSeq) database (http://www.ncbi.nlm.nih.gov/RefSeq/). RefSeq is a public database of nucleotide and protein sequences with corresponding features and bibliographic annotations. The RefSeq database is built and distributed by the NCBI. NCBI builds RefSeq from the sequence data available in the archival database GenBank (Benson et al., 2005). The RefSeq collection is unique in providing a curated, nonredundant, explicitly linked nucleotide and protein database representing significant taxonomic diversity. The RefSeq collection is derived from the primary submissions available in GenBank. GenBank is a redundant archival database that represents sequence information generated at different times, and may represent several alternate views of the protein, names or other information. In contrast, RefSeq represents a nearly non-redundant collection that is a synthesis and summary of available information, and represents the current view of the sequence information, names and other annotations. (Kim et al., 2005).The objective of the present study is to provide a comprehensive phylogenetic and sequence analyses of sperm-surface ADAM genes using the curated and non-redundant RefSeq database. 36 complete refseq CDS (coding sequence) of 9 members of ADAM gene family namely ADAM1,2,3,4,5,6,18,24 and 32 were obtained to investigate its evolution and differentiation within and among species.
2. Materials and Methods
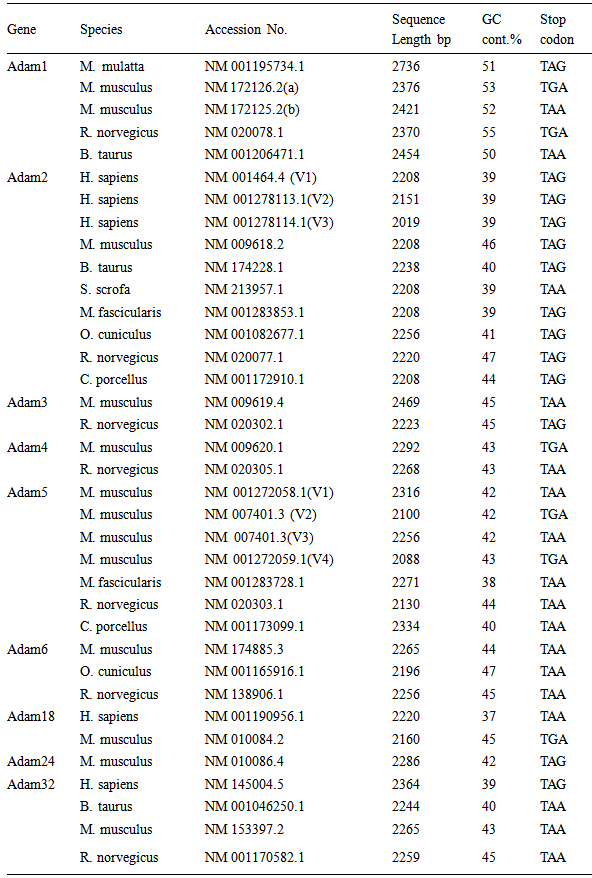
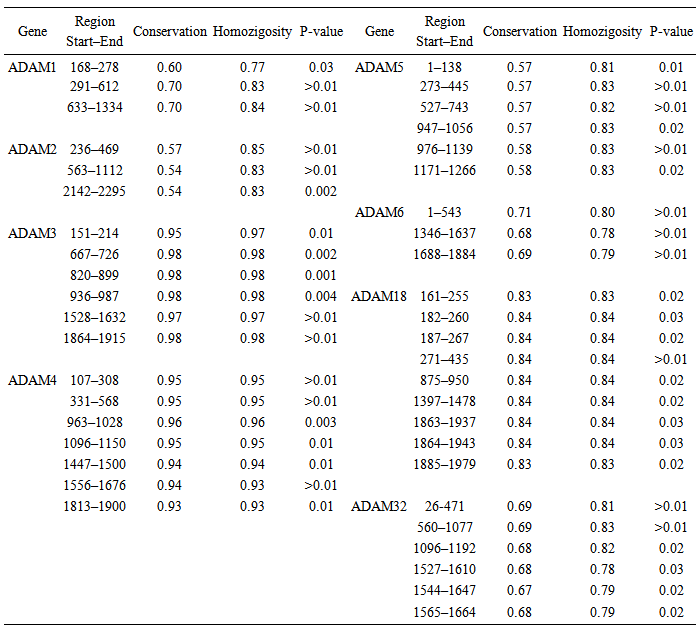
- Data CollectionIn the present study data were obtained from the NCBI Reference Sequence (RefSeq) database (http://www.ncbi.nlm.nih.gov/refseq). RefSeq provides a curated non-redundant sequence database of genomes, transcripts and proteins (Pruitt et al., 2005). The search for ADAM genes sequence were restricted to 9 sperm-surface ADAM genes namely, Adam 1,2,3,4,5,6,18,24 and 32 genes. Only complete, verified, experimentally proofed and non- predictable CDS were considered. The data collection resulted in 36 sequences for the 9 selected genes. Detailed of the 36 genes, species and accession numbers are presented in table (1).
|
3. Results
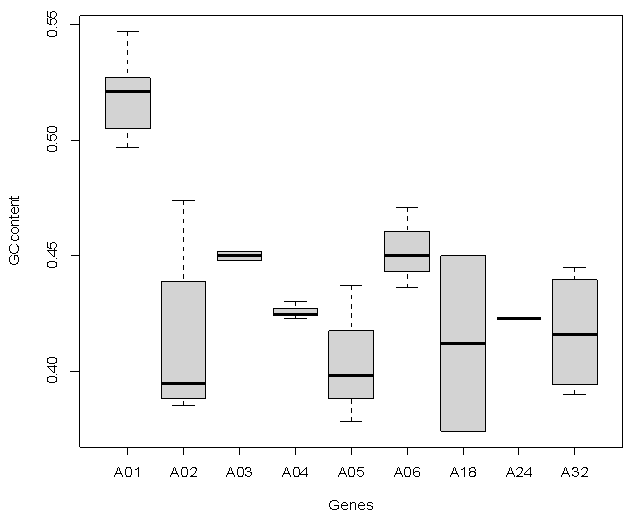
- Sequence Variations & GC-content36 CDS sequences of ADAM1,2,3,4,5,6,18,24 and 32 genes were obtained from NCBI Reference Sequence (RefSeq) database (http://www.ncbi.nlm.nih.gov/refseq). A detailed list of the NCBI refseq accession numbers for the 36 sperm-surface Adam genes as well as percentage GC-content and length of CDS are presented in table (1). The CDS length varied substantially within each of ADAM genes, even for species with close taxonomic relationship (e.g. M. musculus and N. norvegicus) as well as among the 9 genes. The longest sequence was observed in ADAM1, for the 4 species CDS length ranged from 2370 to 2736 bp, where ADAM2 has shortest, and its length ranged form 2019 to 2238 bp. Among the 8 species of ADAM2, homo sapiens had the shortest length for its 3 variants, that is 2019, 2151 and 2208 bp. Figure (1) shows the base composition of the 9 ADAM genes, with exception of ADAM1 no noticeable differences are observed. ADAM1 has the lowest percentage of A and T bases but also ADAM1 has the highest percentages of G and C. These differences are more pronounced when GC-content was considered. The GC-content of all studied sequences ranged from 37 to 55%. Figure (2) shows that the range of variation within each of the 9 genes differs from one gene to another. The range of GC-content% for ADAM 1 was the highest (50–55%) where ADAM2 was the lowest (39–47%). However, ADAM3 and ADAM4 showed exactly similar GC-content, although genes have different CDS length for both of M. musculus and N. norvegicus.
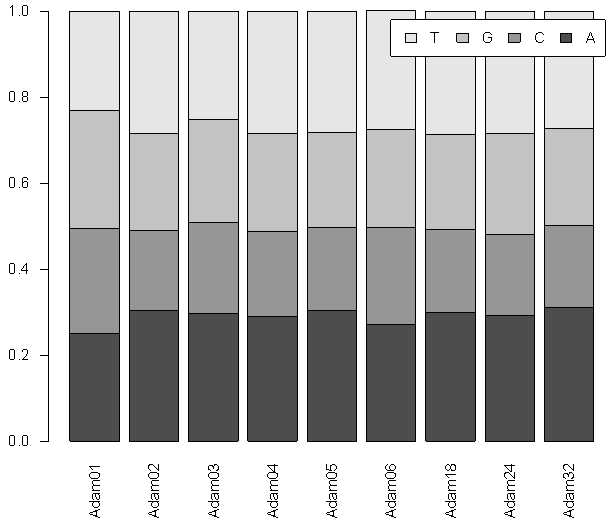 | Figure 1. Plot of the base frequencies of the 36 CDSs for the 9 sperm-surface ADAM genes |
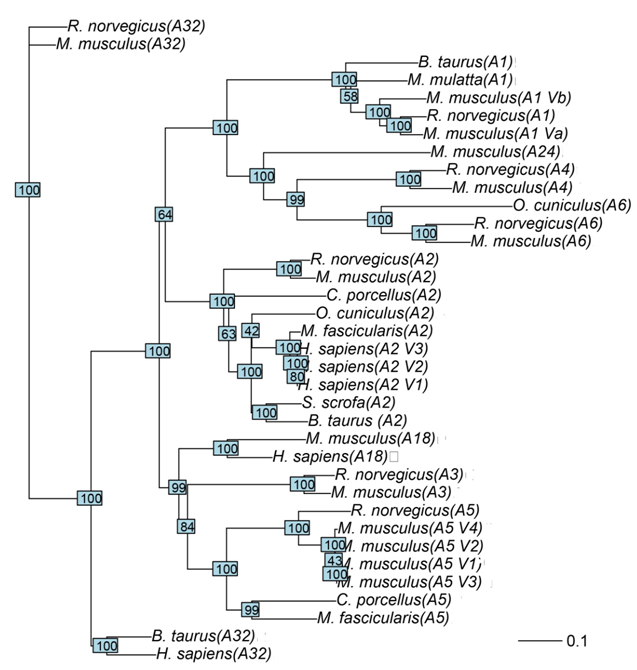 | Figure 3. Neighbor-Joining phylogenetic tree of 9 ADAM genes, the scale bar corresponds to 0.1 substitution per site |
|
|
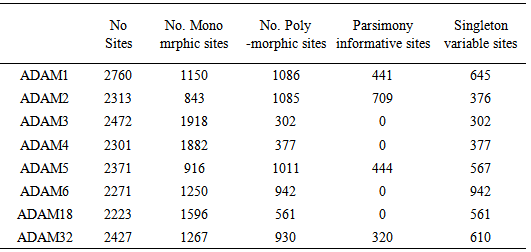
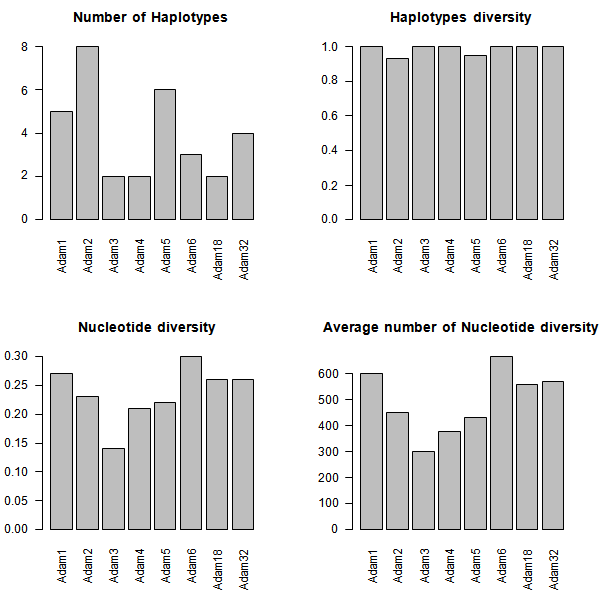 | Figure 4. Measures of DNA polymorphism for Adam genes |
|
4. Discussion
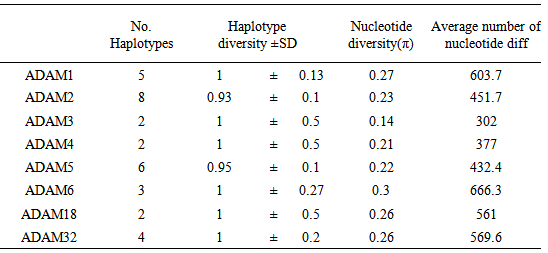
- To the best of our knowledge this is the first time for sperm-surface ADAM genes to be analyzed based on Refseq data. The present study focused on non-redundant and curated RefSeq data containing 36 CDS of 9 sperm-surface ADAM genes.Our results show great variation in the length of CDS sequences of sperm- surface ADAM genes either between or within species for example length variation in variants a and b of ADAM1 in M. musculus. Moreover, variation in stop codon were also observed. These results expand on the previous findings on the molecular evolution of ADAM family genes (Glassey & Civetta, 2004) to show that all ADAM genes involved in male reproductive tract show evidence of being under the selection pressure. Dorus et al. (2010) detected positive selection for ADAM1,2,4,6 and 24 using phylogenetic comparisons among five different species of mammals. Moreover, Finn & Civetta (2010) analyzed 25 members of ADAM gene family and found that all genes expressed in male reproductive tissues showed evidence of positive selection. The same study reported positive selection on codon sites within ADAM1, 2, and 32. This signal of positive selection within ADAM genes might be ascribed to species-specific adaptation of fertilization (Shamsadin et al., 1999, Nishimura et al., 2001 and Yamaguchi et al., 2009).The genomic GC content is one of the key parameters of variation of genome sequences, the value of GC confined to between 25% and 75% (Wu et al., 2012). GC% of the 9 studied ADAM genes ranged between 37 to 55%. This great variation in GC content of sperm-surface ADAM genes is reflection of nucleotide and haplotype diversity (table 3). Choi et al. (2013) reported GC% of 72.9% of non-coding sequences of ADAM2 genes. The length of CDS was shown to be under both functional and structural constrains (Blake, 1983, Blake, 1985, Hawkins, 1988 and Traut, 1988). It is also known that the size distributions of the gene parts (exons, introns, leader and trailer regions, etc.) are under stabilizing selection against extreme lengths (Smith, 1988). Moreover, Oliver & Marin (1996) found that the exon length doesn’t affected by concentration of GC in vertebrate. The authors ascribed this result to lower number of exons in the genes located in the studied regions. Our results (table 1) showed that GC% does not affected by the length of CDS for example; the GC% of three variants of ADAM2 of H. sapeins did not vary with variation of CDS length. As Oliver & Marin (1996) concluded the effect of GC% on the length of CDS might constitute a new evolutionary meaning for compositional variation in DNA GC content.Although ADAM genes are conserved in evolution, not all members of this family are not found in some mammals (Long et al., 2012). In the present study we investigated the phylogenetic of sperm-surface ADAM genes which is important of in understanding the evolutionary history of these genes. We found that the phylogeny of sperm-surface genes was well supported. Our phylogenetic reconstruction supported the overall orthology of sperm-surface ADAM genes. ADAM1,4,6 and 24 were grouped together in one clade. This clade did not include H. sapiens which indicated these genes do not have human orthologues. This result is in agreement with the finding of Choi et al. (2004). However, the only differences in the topology of tree between our results and the outputs of both Finn & Civetta (2010) and Grayson & Civetta (2013) is inclusion of ADAM32 of all species in the same clade. Finn & Civetta (2010) used redundant data from several species where Finn & Civetta (2010) worked only on Mus species. In fact these results shade the light into the significance of examining different factors such as selection within specific groups or clades, because the effect of selection might be impaired by phylogenetic analysis that include various species. The same conclusion was drawn by Civetta (2012) and Garyson & Civetta (2013).Our results extend to previous finding that considerable sequence diver- sity exists among sperm-surface ADAM genes. We found that this was a reflected by the nucleotide diversity and the average number of nucleotide differences. Our analysis revealed that individual members of sperm-surface ADAM genes differs widely in their average number of haplotypes, nucleotide diversity and the average number of nucleotide differences. Moreover, not all polymorphic sites were informative where, ADAM3,4,6 and 18 were lacking informative sites.The analysis of the conserved regions of sperm-surface ADAM genes sup-ported the orthology within each member of this gene family. This conclusion is justifiable with the high estimates of conservation, homozigosity and P-values. The conservation within each of ADAM genes is in a good agreement with their essential functionality in vivo as determined by knocked out mice. In conclusion, the in silico analysis of the 9 sperm-surface ADAM genes showed a great deal of variation among and within this genes indicating the presence of localized signals of selection pressure on these genes. Moreover, ADAM1,4,6 and 24 have no human orthologues. More importantly, this result suggests that ADAM1,2,3,4,5,6, 18,24 and ADAM32 of B.taurus and H. sapiens were descendent form one ancient ancestor, where ADAM32 of R. norvegicus and M. musculus have another ancestor.
 Abstract
Abstract Reference
Reference Full-Text PDF
Full-Text PDF Full-text HTML
Full-text HTML