-
Paper Information
- Next Paper
- Previous Paper
- Paper Submission
-
Journal Information
- About This Journal
- Editorial Board
- Current Issue
- Archive
- Author Guidelines
- Contact Us
American Journal of Medicine and Medical Sciences
p-ISSN: 2165-901X e-ISSN: 2165-9036
2026; 16(3): 898-900
doi:10.5923/j.ajmms.20261603.10
Received: Feb. 2, 2026; Accepted: Feb. 26, 2026; Published: Mar. 4, 2026

Improving Early Diagnosis and Prognosis of Chronic Hepatitis C Based on the Assessment of IL-10 (G108A) and TNF-α (G308A) Gene Polymorphisms
Khayrullaeva D. Kh.1, Yuldasheva D. Kh.2
1Tashkent State Medical University, Tashkent, Uzbekistan
2Bukhara State Medical Institute, Bukhara, Uzbekistan
Copyright © 2026 The Author(s). Published by Scientific & Academic Publishing.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/

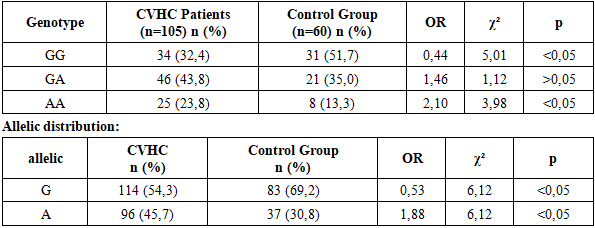
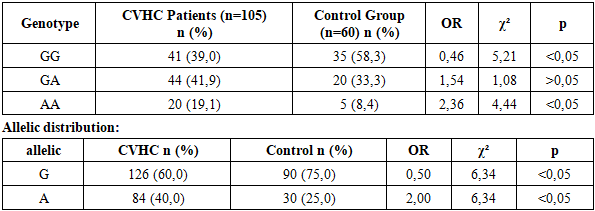
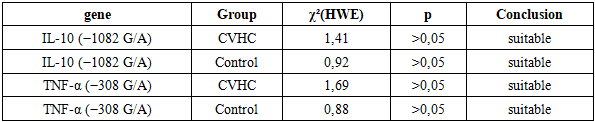
This article demonstrates that IL-10 (−1082 G/A) and TNF-α (−308 G/A) gene polymorphisms are significantly associated with susceptibility to chronic viral hepatitis C and its progression. The analysis showed that for both genes, the AA genotype and A allele were significantly more frequent in patients compared to the control group, confirming their role as risk factors for the development of chronic viral hepatitis C. Conversely, the GG genotype and G allele were more common in the control group, indicating their protective effect against the disease. Additionally, the genotype distributions of IL-10 and TNF-α in both patients and controls were in accordance with Hardy–Weinberg equilibrium, supporting the genetic representativeness of the study sample.
Keywords: Chronic viral hepatitis C, Gene, Polymorphism, Genotype, Disease
Cite this paper: Khayrullaeva D. Kh., Yuldasheva D. Kh., Improving Early Diagnosis and Prognosis of Chronic Hepatitis C Based on the Assessment of IL-10 (G108A) and TNF-α (G308A) Gene Polymorphisms, American Journal of Medicine and Medical Sciences, Vol. 16 No. 3, 2026, pp. 898-900. doi: 10.5923/j.ajmms.20261603.10.
1. Introduction
- Chronic viral hepatitis C (CVHC) is a widespread infectious disease worldwide and in Uzbekistan, causing significant social and economic burden. According to the World Health Organization, a large proportion of individuals infected with the hepatitis C virus (HCV) progress to a chronic form, which over 10–20 years may lead to liver cirrhosis, hepatocellular carcinoma, and severe hepatic dysfunction. In addition, HCV infection has long-term effects on patients’ health and results in socio-economic consequences such as loss of work capacity, increased costs of medications, and expenses related to inpatient treatment. The clinical course of CVHC and the development of its complications are not limited to virological factors alone but are also closely associated with the patient’s immune response and genetic characteristics. In particular, genetic polymorphisms regulating cytokine synthesis play an important role in determining the activity of inflammation, the rate of fibrosis progression, and disease prognosis. In this regard, IL-10 (G108A) and TNF-α (G308A) gene polymorphisms are considered key genetic markers of scientific and practical significance in the pathogenesis of chronic hepatitis C [1,3,4]. Studies by Powell et al. (2000) demonstrated that the TNF-α (G308A) gene polymorphism is associated with higher inflammatory activity and accelerated development of fibrosis in liver tissue. The authors emphasized that patients carrying the A allele of the TNF-α gene have a more severe disease course and a higher risk of complications. In CVHC, cytokines such as IL-10 and TNF-α play a central role in regulating immune responses. IL-10 is an anti-inflammatory cytokine that suppresses intercellular inflammation and immune reactions. The G108A polymorphism of the IL-10 gene may influence IL-10 production and its functional activity. The G308A polymorphism of the TNF-α gene can affect cytokine levels and the intensity of inflammation, thereby increasing the risk of liver tissue damage, fibrosis, and progression to cirrhosis. In clinical practice, the diagnosis of chronic hepatitis C is most often based on clinical findings and standard laboratory parameters, which limits the possibilities for early detection of the disease and prediction of complications. In contrast, the assessment of genetic polymorphisms makes it possible to identify the underlying mechanisms influencing the clinical course of the disease, to detect high-risk patients at an early stage, and to implement individualized monitoring strategies [2,5]. Taking these aspects into account, the aim of the present study is to evaluate the key genetic polymorphisms that determine the clinical course and progression of chronic hepatitis C in affected patients, to identify their associations with laboratory and clinical parameters, and, based on the obtained data, to improve early diagnosis and disease prognosis.
2. Materials and Methods
- In this study, the associations of genetic polymorphisms (IL-10 (G108A) and TNF-α (G308A)) with clinical, laboratory, and virological parameters were evaluated. The main group included 105 patients with a confirmed diagnosis of chronic hepatitis C, of whom 58 were male and 47 were female. The patients’ ages ranged from 25 to 76 years, with a mean age of 49.6 ± 1.2 years. The control group consisted of 60 healthy individuals with no history of hepatitis C infection. They were matched to the main group by sex and age: 32 males and 28 females, aged 20–63 years (mean age 41.9 ± 1.1 years). The study was conducted with approval from the Ethics Committee of Bukhara Medical Institute. Genetic analysis. DNA was extracted from peripheral blood using standard methods. IL-10 (G108A) and TNF-α (G308A) polymorphisms were determined using the PCR–RFLP method. Genotypes were classified as GG, GA, and AA variants. Comparative analysis of qualitative variables was performed using the χ² (chi-square) test or Fisher’s exact test. Genotype and allele frequency distributions were calculated, and odds ratios (OR) and relative risks (RR) with 95% confidence intervals were determined. Associations between genetic polymorphisms, clinical features, and laboratory parameters were assessed using Spearman’s or Pearson’s correlation coefficients. Logistic regression analysis was performed to predict complications of chronic hepatitis C. Statistical significance was defined as p < 0.05.
3. Results and Analysis
- Genetic analyses were performed on DNA samples isolated from peripheral blood. IL-10 (G108A) and TNF-α (G308A) gene polymorphisms were identified using the PCR–RFLP method. Genotypes were classified into GG, GA, and AA variants. The distribution of genotypes was evaluated for compliance with Hardy–Weinberg equilibrium.
|
|
|
 Abstract
Abstract Reference
Reference Full-Text PDF
Full-Text PDF Full-text HTML
Full-text HTML