-
Paper Information
- Next Paper
- Previous Paper
- Paper Submission
-
Journal Information
- About This Journal
- Editorial Board
- Current Issue
- Archive
- Author Guidelines
- Contact Us
American Journal of Medicine and Medical Sciences
p-ISSN: 2165-901X e-ISSN: 2165-9036
2026; 16(3): 887-891
doi:10.5923/j.ajmms.20261603.08
Received: Feb. 12, 2026; Accepted: Mar. 2, 2026; Published: Mar. 4, 2026

Focality and Heterogeneity of Cervical Intraepithelial Neoplasia Grades I–II: The Role of Integrated Assessment and FISH Analysis
Kunduz A. Aхmedova1, Dilfuza A. Alieva2, Natalia S. Nadyrkhanova3
1Independent Researcher, Republican Specialized Scientific and Practical Medical Center for Maternal and Child Health, Tashkent, Uzbekistan
2MD, PhD, Professor, Head of the Scientific Advisory Outpatient Clinic “Family and Marriage”, Republican Specialized Scientific and Practical Medical Center for Maternal and Child Health, Tashkent, Uzbekistan
3MD, PhD, Director of the Republican Specialized Scientific and Practical Medical Center for Maternal and Child Health, Tashkent, Uzbekistan
Copyright © 2026 The Author(s). Published by Scientific & Academic Publishing.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/

Early detection of neoplastic transformation of the cervical epithelium remains a key challenge in modern gynecology and cancer prevention. The aim of this study was to evaluate the diagnostic and prognostic potential of fluorescence in situ hybridization (FISH) for the detection of early molecular genetic alterations in the cervical epithelium in patients with cervical intraepithelial neoplasia (CIN). The study included 70 women aged 30–65 years with CIN I–II and invasive cervical cancer. All patients underwent a comprehensive examination including liquid-based cytology, high-risk human papillomavirus (HR-HPV) testing, and FISH analysis using locus-specific DNA probes targeting the p16 (CDKN2A), TP53, and ATM genes. Statistical analysis included calculation of p-values, 95% confidence intervals, correlation analysis, and binary logistic regression. The frequency of genetic aberrations detected by FISH increased significantly with CIN progression: 44.0% in CIN I, 58.3% in CIN II, and 57.1% in cervical cancer (p = 0.03). A moderate positive correlation was found between CIN grade and FISH positivity (r = 0.46; p = 0.002). ATM gene amplification was identified in 36% of cases and was associated with an increased risk of neoplastic progression. A threshold of ≥10% of epithelial cells with abnormal FISH signals was considered clinically significant. FISH demonstrated high diagnostic and prognostic value and may be recommended as an adjunct method for risk stratification and individualized management of patients with CIN.
Keywords: Human papillomavirus, Cervical intraepithelial neoplasia, FISH, Liquid-based cytology, p16, TP53, ATM, Genetic aberrations
Cite this paper: Kunduz A. Aхmedova, Dilfuza A. Alieva, Natalia S. Nadyrkhanova, Focality and Heterogeneity of Cervical Intraepithelial Neoplasia Grades I–II: The Role of Integrated Assessment and FISH Analysis, American Journal of Medicine and Medical Sciences, Vol. 16 No. 3, 2026, pp. 887-891. doi: 10.5923/j.ajmms.20261603.08.
1. Introduction
- Cervical cancer (CC) is one of the most common malignancies of the female reproductive system, particularly prevalent in developing countries, mainly due to limited access to effective diagnostic tests [1].It is well known that HPV-associated cervical carcinogenesis is a multifactorial, progressive disease that usually passes through several stages of intraepithelial precancerous lesions of the cervix. In approximately 10% of cases, low-grade cervical intraepithelial neoplasia (CIN I) may progress to high-grade CIN (CIN II–III) and cervical cancer; therefore, early identification of cases with potential malignant and premalignant progression is critically important.Cytological screening programs are widely used for the diagnosis of cervical cancer. Cytological examination of cervical epithelial cells (Pap smears, liquid-based cytology) allows detection of early dysplastic changes, making it possible to implement preventive measures against malignant transformation. However, several studies have demonstrated that up to 50% of precancerous cervical lesions may be missed due to false-negative Pap test results [2,3]. Moreover, cytological examination does not assess the risk of disease progression or clearance of HPV infection leading to regression of pathological changes.In this context, molecular genetic methods, including FISH, are of particular interest, as they enable detection of early structural genomic alterations and provide biologically grounded risk stratification for CIN progression in clinical practice [4,5].Carcinogenesis is known to be driven by genetic modifications. Alterations in proto-oncogenes or tumor suppressor genes lead to changes in gene expression and, consequently, in the levels of functionally active protein molecules. These genetic disturbances ultimately result in impaired differentiation and maturation of cervical squamous epithelial cells, loss of cell cycle control, and the development of uncontrolled cellular proliferation [6–8].To date, a number of genes exhibiting different expression levels in normal epithelium, intraepithelial dysplasia of varying severity, and invasive cervical cancer have been identified. Thus, M. Del Pino et al. demonstrated a relationship between the expression level of the Cyclin-dependent kinase inhibitor 2A gene (CDKN2A, p16) and the severity of cervical lesions, with method sensitivity reaching 81% in HSIL (CIN III) [9]. Among genetic alterations detected in epithelial cells in cervical dysplasia and cervical cancer, the key role of the chromosomal region 9p21.3, where the CDKN2A (p16) gene is located, has been confirmed [10].The p16 gene product is involved in cell cycle regulation by inducing arrest at the G1 and G2/M checkpoints and acts as a negative regulator of cell proliferation [11,12]. Genetic alterations of p16 result in reduced CDKN2A protein expression. In neoplastic processes, p16 inactivation may be caused by promoter methylation, point mutations, or homozygous deletion [13].One of the methods for detecting genetic alterations associated with cervical epithelial cell transformation is fluorescence in situ hybridization (FISH), performed on cytological or histological specimens. FISH analysis using locus-specific DNA probes allows identification of gene rearrangements, as well as deletions and amplifications of chromosomal regions containing clinically significant markers, including 9p21.3 (p16) and 17p13.1 (TP53).At the same time, assessment of p16, TP53, ATM, and other molecular markers of cervical neoplasia development is associated with several limitations due to tumor tissue heterogeneity, the need for invasive biopsy, and differences in sensitivity and specificity of the applied methods [13,14]. Therefore, there remains a need for diagnostic approaches that combine high accuracy with practical applicability in clinical settings.Aim of the StudyTo evaluate the capabilities of the FISH method in detecting early genetic rearrangements of the cervical epithelium and their clinical significance in CIN progression.
2. Materials and Methods
- Study PopulationThe study included 70 patients aged 30–65 years, divided into the following groups:• CIN I — 25 patients (14 HPV-positive, 11 HPV-negative);• CIN II — 24 patients (14 HPV-positive, 10 HPV-negative);• Comparison group (cervical cancer) — 21 patients (19 HPV-positive, 2 HPV-negative), included prior to initiation of specialized treatment.The median age was 36 (34–38) years in the CIN I group, 42 (40–44) years in the CIN II group, and was significantly higher in patients with invasive cervical cancer — 52 (50–54) years. Age differences between groups were statistically significant (p < 0.001), reflecting an age-associated increase in neoplastic severity.Inclusion Criteria (CIN I–II Groups)1. Women aged 30–65 years;2. Presence or absence of high-risk HPV (considered in analysis);3. Cytological diagnosis of LSIL or ASC-H, HSIL according to the Bethesda system;4. Regular menstrual cycle;5. Ability to comply with the study protocol;6. Signed informed consent.Inclusion Criteria (Cervical Cancer Group)1. Histologically confirmed invasive cervical cancer;2. No prior specialized treatment;3. Signed informed consent.Exclusion Criteria1. Pregnancy;2. Postpartum period and lactation;3. Hormonal therapy;4. Acute inflammatory diseases of specific or nonspecific etiology;5. Decompensated renal, hepatic, or pulmonary diseases;6. Neuropsychiatric disorders;7. Inability to comply with the study protocol.CIN Group Formation CriteriaCIN I groupAt the initial stage (prior to histological verification), inclusion was based on a cytological diagnosis of LSIL according to the Bethesda system.CIN II groupInclusion criterion was a cytological diagnosis of ASC-H or HSIL according to the Bethesda system.Due to the stepwise nature of diagnosis and follow-up, group formation was dynamic. Patients initially diagnosed with ASC-H who were histologically confirmed as CIN I after targeted cervical biopsy were assigned to the CIN I group. Final group allocation was based on histological findings.Follow-UpPatients with CIN I were followed for 24 months. Follow-up examinations, including liquid-based cytology, HPV testing, and extended colposcopy, were performed every 6 months. In cases of worsening clinical, cytological, or colposcopic findings, targeted cervical biopsy with subsequent histological examination was performed.Patients with CIN II underwent targeted cervical biopsy as part of the standard diagnostic algorithm for morphological verification and management decision-making.Ethical ConsiderationsAll patients provided written informed consent. The study was approved by the Bioethics Committee (protocol dated June 12, 2024).Statistical AnalysisStatistical analysis was performed using IBM SPSS Statistics for Windows, version 26.0 (IBM Corp., Armonk, NY, USA). Quantitative variables are presented as the median and interquartile range, Me (Q1–Q3). Intergroup comparisons of quantitative variables were performed using the Mann–Whitney U test.Categorical variables were analyzed using Fisher’s exact test. Event frequencies were reported with 95% confidence intervals (95% CI).The relationship between CIN grade (CIN I, CIN II, cervical cancer) and FISH-detected genetic aberrations was assessed using Spearman’s rank correlation coefficient.Binary logistic regression analysis was applied to evaluate the influence of CIN grade and HPV status on the probability of detecting genetic aberrations. Differences were considered statistically significant at p < 0.05.MethodsLiquid-based cytology was performed prior to cervical biopsy. FISH analysis was carried out using LSP p16/Cep9 and LSP TP53/ATM DNA probes. At least 100 interphase nuclei were analyzed in each sample. Signal interpretation was performed in accordance with the International System for Human Cytogenomic Nomenclature (ISCN).Quality Control, Reproducibility, and ValidationA multilevel quality control system was applied. Centromeric probes (Cep9) served as internal controls. Morphologically unchanged epithelial cells were used as negative controls, while specimens with previously confirmed genetic aberrations served as positive controls.Reproducibility was assessed through repeated analysis of selected samples by independent observers; discrepancies did not affect final classification. Validation was performed by comparing FISH results with cytological and histological findings.Clinical Significance ThresholdA ≥10% proportion of epithelial cells with abnormal FISH signals was considered clinically significant. Values below 10% were regarded as borderline and were not used for clinical decision-making.FISH Sample Preparation TechniqueThe FISH sample preparation technique for interphase nuclei of cervical epithelial cells was developed based on a direct fixation method used in cytogenetic studies. Hybridization and analysis of DNA probe signals for p16, TP53, and ATM genes were performed in 70 patients with CIN I, CIN II, and invasive cervical cancer.Biological material consisted of cells obtained from the stratified squamous epithelium of the cervix and cervical canal and transferred into a liquid-based cytology fixative (YES PATH LBC Solution, Gyn 17 ml, Republic of Korea).To assess the quality and suitability of the developed preparation method for FISH analysis, additional cytological slides prepared from the same cellular fixative were stained with Giemsa. Subsequent FISH signal analysis demonstrated partial disruption of cell membranes and loosening of cytoplasmic structure, reducing intracellular density. This facilitated a decrease in non-hybridized probe background fluorescence, improving signal visualization. Nuclear integrity, clear contours, and uniform distribution without overlap were preserved, minimizing interpretation errors and enabling accurate copy number assessment. At least 100 interphase nuclei were analyzed per sample in accordance with ISCN recommendations.DNA Probe CharacteristicsLSP p16/Cep9 DNA probe detects the CDKN2A (p16) gene and chromosome 9 copy number. The green centromeric signal corresponds to locus 9p13.2, while the red signal marks 9p21.3. Normally, two green and two red signals are observed. Signal variations indicate trisomy, monosomy, monoallelic or biallelic deletion, or amplification/rearrangement of p16. LSP TP53/ATM DNA probe detects rearrangements involving TP53 (17p13.1) and ATM (11q22.3). Signal number changes reflect deletions, amplifications, chromosomal gains or losses, and gene rearrangements.
3. Results
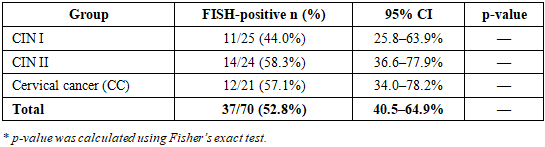
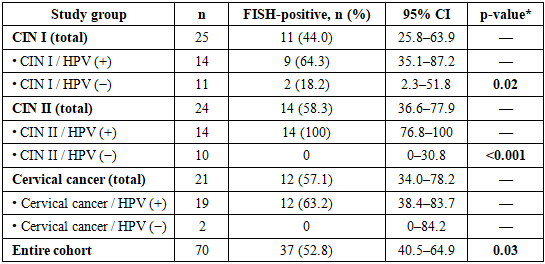
- Genetic aberrations detected by FISH were identified in 37 of 70 patients (52.8%; 95% CI: 40.5–64.9). The frequency of FISH-positive cases increased significantly with CIN progression (p = 0.03). ATM gene amplification was detected in 36% of patients, predominantly in CIN II and cervical cancer. A moderate positive correlation between CIN grade and genetic aberration frequency was observed (r = 0.46; p = 0.002).Logistic regression analysis demonstrated that CIN grade (OR = 2.1; p = 0.01) and high-risk HPV status (OR = 2.8; p = 0.004) were independent predictors of FISH positivity.
|
|
4. Discussion
- Detection of ATM gene amplification reflects activation of DNA damage response mechanisms and indicates pronounced genomic instability. This abnormality may serve as a marker of unfavorable prognosis and increased risk of CIN progression.FISH analysis with assessment of the proportion of genetically altered cells enables a shift from a purely morphological approach to biologically oriented risk stratification. Detection of ≥10% of cells with abnormal signals justifies shorter follow-up intervals and earlier implementation of invasive diagnostic or therapeutic interventions.
5. Conclusions
- 1. The frequency of genetic aberrations detected by FISH increases with CIN progression.2. ATM gene amplification may be considered a marker of increased risk of neoplastic progression.3. A clinically significant FISH positivity threshold is ≥10% of cells with abnormal signals.4. FISH is advisable for individualized management of patients with CIN.5. Cervical screening in CIN patients, including Pap smear cytology, liquid-based cytology, and HR-HPV PCR testing, should be complemented by FISH analysis using LSP probes targeting oncogenic risk markers, including TP53, p16, and ATM gene rearrangements.
 Abstract
Abstract Reference
Reference Full-Text PDF
Full-Text PDF Full-text HTML
Full-text HTML