-
Paper Information
- Next Paper
- Paper Submission
-
Journal Information
- About This Journal
- Editorial Board
- Current Issue
- Archive
- Author Guidelines
- Contact Us
American Journal of Biochemistry
p-ISSN: 2163-3010 e-ISSN: 2163-3029
2017; 7(5): 100-113
doi:10.5923/j.ajb.20170705.03

A Novel Thermotolerant β-glucosidase from Aspergillus nidulans has Activity across a Broad pH Profile and a Likely Bacterial Origin
Richard Auta1, 2, Iza Radecka1, Paul Hooley1
1Faculty of Science and Engineering, University of Wolverhampton, UK
2Department of Biochemistry, Kaduna State University, Kaduna, Nigeria
Correspondence to: Richard Auta, Faculty of Science and Engineering, University of Wolverhampton, UK.
| Email: |  |
Copyright © 2017 Scientific & Academic Publishing. All Rights Reserved.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/

This study reports the purification and characterization of a recombinant β-glucosidase expressed from an AOX1 promoter in Pichia pastoris carrying an Aspergillus nidulans (A. nidulans) cloned gene. β-glucosidase was optimally active at 50°C and pH 5.5, though it had a broad pH range of pH 3.0 – 10.0 and a broad temperature range of 10 – 80°C. β-glucosidase showed very high affinity to para-Nitrophenyl β-D-glucopyranoside (pNPG). Evidence for a bacterial origin of this gene (AN1804) was provided by the absence of introns, absence of some fungal specific amino acid insertions in its encoded protein sequence, automatic annotation as “periplasmic” and unusual positions in phylogenetic trees showing similarities to bacterial proteins.
Keywords: β-glucosidase, A. nidulans, Horizontal gene transfer, Para-Nitrophenyl β-D-glucopyranoside
Cite this paper: Richard Auta, Iza Radecka, Paul Hooley, A Novel Thermotolerant β-glucosidase from Aspergillus nidulans has Activity across a Broad pH Profile and a Likely Bacterial Origin, American Journal of Biochemistry, Vol. 7 No. 5, 2017, pp. 100-113. doi: 10.5923/j.ajb.20170705.03.
Article Outline
1. Introduction
- The conversion of cellulolytic materials into fermentable sugars requires the action of endoglucanase, exoglucanase and β-glucosidase which work in concert to hydrolyse cellulose [1-3]. Endoglucanases attack amorphous sites of the cellulose, randomly yielding oligosaccharides of various lengths, while exoglucanases attack crystalline cellulose from both reducing and non-reducing ends generating glucose or cellobiose as major products [4]. Both endoglucanases and exoglucanases hydrolyse β-1, 4-glycosidic bonds. β-glucosidases finally hydrolyse cellobiose and other oligosaccharides into glucose. The catalysis of cellobiose is important since the accumulation of cellobiose creates feed-back inhibition. β-glucosidase is generally responsible for the regulation of the whole cellulolytic process and is a rate limiting factor during enzymatic hydrolysis of cellulose, as both endoglucanase and exoglucanase activities are often inhibited by cellobiose [5]. β-glucosidases (also known as cellobiase, β-D-glucosidase, β-glucoside glucohydrolase, p-nitrophenyl β-glucosidase, and β-1,6-glucosidase; EC 3.2.1.21) catalyze the hydrolysis of terminal non-reducing residues in β-D-glucosides by the transfer of a glycosyl group between oxygen nucleophiles to release glucose [6]. β-glucosidases are well distributed in nature and they play important roles in a large number of biological processes [7]. In humans for instance, β-glucosidase is involved in the hydrolysis of glucosyl ceramides [8]. In plants, β-glucosidase is involved in the synthesis of β-glucan during cell development, defense mechanisms, fruit ripening and pigment metabolism while in microorganisms it is involved in cellulase induction and cellulose hydrolysis [9-11]. Aspergillus niger has been identified as one of the most efficient producers of β-glucosidase [12, 13]. Some of the enzyme properties considered important for production, purification and its application are its molecular weight, optimal pH and temperature for enzyme activity and stability [14]. Different isolates of Aspergillus have been reported to express β-glucosidases with differences in catalytic activity, thermostability and optimum pH [12, 15]. High temperature and low pH tolerant enzymes are preferred for the hydrolysis of lignocellulolytic materials because most current pretreatment strategies depend on acid and heat [16]. The recently completed genome projects for the genus Aspergillus have provided a rich resource for the identification of β-glucosidases with novel properties [17]. Among the first published Aspergillus genome sequences from a group of related filamentous fungi are the genome sequences of A. nidulans, A. oryzae, A. niger, A. fumigatus, A. terreus, A. clavatus, A. parasiticus and Neosartorya fischeri [18-20]. Galagan et al. [21] reported the genome sequence of the model organism Aspergillus nidulans by comparative analysis with A. fumigatus and A. oryzae. Coutinho et al. [22] reported three GH1 β-glucosidases and twenty one GH3 β-glucosidases from A. nidulans FGSC A4. Manual verification of their results indicated that 28.4% of the A. nidulans ORF’s do not contain a secretion signal, of which 40% may be secreted through a non-classical method. Though there is no single well defined method for the classification of β-glucosidase, four methods for their classification appear in the literature. β-glucosidases are classified on the basis of substrate specificity, encoding gene nucleotide sequence identity, mechanism of action and localization within the cell [23-25]. Based on substrate specificity, β-glucosidases have conventionally been divided into cellobiase (with specificity for cellobiose), aryl-β-glucosidase (with specificity for p-nitrophenyl-β-D-glucopyranoside, pNPG) and broad specificity β-glucosidases [23]. CAZy classification of Glycosyl hydrolases (GH) has placed β-glucosidase in family GH1 and GH3 based on their amino acid sequences but they can also be found in GH family 5, 9, 17, 30 and 31 [24, 26-28]. Coutinho et al. [22] also described A. nidulans, A. niger and A. oryzae to have 24, 20 and 26 GH1/GH3 enzymes respectively for each species. Most organisms such as Aspergillus selectively use substrates from a mixture of different carbon sources, but the presence of glucose which is the most preferred carbon source represses the expression of β-glucosidase and the activity of catabolic system that enable the degradation of cellulose [29-31]. Hence, the regulated expression of high titres of enzymes may require heterologous expression in alternative hosts using inducible promoters. Protein expression systems include Escherichia coli (E. coli), yeast; baculovirus infected insect cells and mammalian cells [32]. Saccharomyces cerevisiae (S. cerevisiae) and the methylotrophic yeast Pichia pastoris (P. pastoris) are the commonly used yeast strains for protein expression with commercial cloning and expression kits available [33]. For example, Bauer et al. [34] have already reported the successful cloning and expression of a number of plant and fungal cellulases using a Pichia pastoris expression vector with a methanol inducible AOX1 promoter. They reported the production of five putative β-glucosidase enzymes from A. nidulans with the accession numbers AN227.2, AN2612.2, AN0712.2, AN1551.2 and AN1804.2. During the course of a bioinformatics based study of candidate β-glucosidases, a number of A. nidulans accessions with curious characteristics were noted. The aim of this paper is to characterise novel β-glucosidase enzymes of likely bacterial origin from A. nidulans so as to suggest possible selective advantages that explain their retention by the fungal host.
2. Methods
2.1. Keyword Searches
- The Aspergillus comparative database [35] was initially searched using the keyword “β-glucosidase”. A BLASTp, Basic Local Alignment Search Tool (peptide), search (using default database parameters) in the Aspergillus comparative database was then conducted against each of the resulting keyword FASTA sequences. Signal sequence peptide presence was checked using SignalP 4.0 server. Once identified, signal sequences were manually removed from an amino acid sequence at the specified cleavage site. BLASTp search was again conducted to identify more possible β-glucosidases, as previous searches may have been biased by the similarities within the signal sequence alone. A keyword search was also conducted on the Uniprot and the FungiDB database using the text search term “β-glucosidase”. Individual accessions from fungi were then collated. An updated list of accession numbers for the β-glucosidases was then checked individually using a BLASTp search on the Aspergillus comparative database. The results were then checked for amino acid length, number of introns, signal peptide, GH family and putative matches. Multiple sequence alignment was performed with a TCoffee (Tree-based Consistency Objective Function For Alignment Evaluation [36] to identify regions of similarity, structure and function between sequences. BLAST (Basic Local Alignment Search Tool) [37] was used in comparing primary biological sequence information, such as the amino-acid sequences of different proteins or the nucleotides of DNA sequences. Treeview [38] was used for displaying phylogenies. Phylogenetic trees (neighbour joining) of the GH1 and GH3 β-glucosidase sequences were constructed using ClustalX and Treeview [38, 39] with the bootstrap value set at 2000. A value of 2000 bootstraps was chosen to calculate the robustness of each branch, indicating the number of attempts to draw a particular tree. SignalP 4.0 [40] was used in the identification of signal peptides and prediction of their cleavage sites.
2.2. Small Scale Cultivation of the Recombinant P. pastoris β-glucosidase Clone
- The experiment was carried out to determine protein production yield and activity of the AN1804 clone [34] a putative β-glucosidase enzyme from A. nidulans expressed in a Pichia clone using the pPICZ expression system [34] (deposited in the Fungal Genetic Stock Centre [FGSC] USA). The engineered AN1804 clone was inoculated into 25 ml of Buffered Glycerol-Complex Medium (BMGY) in a 250 ml flask. The cell cultures were incubated on an orbital shaker at 30°C and 250 rpm for 16 - 18 hours to grow in the repressive medium before induction. When the culture concentration reached an OD600 = 3, the cells were harvested by centrifuging at room temperature and 5000 rpm for 5 minutes. The cell pellet was re-suspended in 90 ml Buffered Methanol Complex Medium (BMMY - Yeast extract 1%, Peptone 2%, 100 mM potassium phosphate (pH 6.0), YNB 1.34%, Biotin 4 x 10-5%, Methanol 0.5%) medium until OD6oo = 1 to induce expression. The culture was incubated in an orbital shaker at 150 rpm and 30°C for 4 days. The growth and induction took place in 1 litre flasks and the cell culture was induced every 24 hour after the start of cultivation with methanol to 0.5% of the total volume of the culture to compensate for methanol loss caused by evaporation from medium and uptake by cells. Crude enzyme samples (1.5 ml) were withdrawn from the culture broth after 0, 6, 12, 24, 48, 72 and 96 hours and filter sterilized using syringe filter (0.22 µm) to separate mycelial cells from culture filtrates. Assays were conducted to determine enzyme activity using p-nitrophenyl- β-D-glucopyranoside (pNPG) as substrate. The protein concentration was determined using the Bradford method [41].
2.2.1. Crude Enzyme Production
- The fermentation broth was centrifuged at 8000 rpm for 10 minutes to remove the cells, and the crude β–glucosidase in the culture supernatant was precipitated by adding solid ammonium sulphate (NH4)2SO4 at 60% saturation. The solution was kept overnight at 0 – 4°C. The precipitated proteins were collected by centrifugation at 26000 rpm for 20 minutes. The resultant pellet was re-dissolved in approximately 20 ml of 20 mM phosphate buffer (pH 7.0) and dialyzed against phosphate buffer for 2 days with 3 changes of buffer. The insoluble proteins were removed by centrifugation at 8000 rpm for 10 minutes.The dialysis tubing (Sigma-Aldrich D-9777) with average flat width of 25 mm was initially treated by washing in running water for 3 – 4 hours to remove glycerin. This was followed by treatment with a 0.3% sodium sulfide solution at 80°C for 1 minute to remove sulfur compounds. The tube was then washed with hot water (60°C) for 2 minutes followed by acidification with 0.2% sulfuric acid. Finally, the dialysis tubing was thoroughly washed with hot water to remove the acid following the manufacturer’s instructions.
2.2.2. Enzyme Assay
- β-glucosidase catalyses the hydrolysis of p-nitrophenol-β-glucoside to p-nitrophenol and glucose. Under alkaline conditions, the p-nitrophenolate anion absorbs light at 400 - 450 nm. The amount of enzyme present is therefore determined by measuring the amount of p-nitrophenolate anion produced in the reaction. β-Glucosidase was assayed using p-nitrophenyl- β-Dglucopyranoside (pNPG) as substrate as specified in Ghose and Bisaria [42]. Appropriately diluted enzyme samples of 25 µl were incubated with 25 µl of 10 mM pNPG in citrate buffer (0.05 M, pH 4.8) and 50 µl of citrate buffer (0.05 M, pH 4.8) at 40°C for 15 minutes. The reaction was terminated by adding 100 µl of 0.2 M Na2CO3 solution. Appropriate blanks devoid of enzyme or substrate were also run in parallel to the enzyme assay. The colour developed due to liberation of p-Nitrophenol (pNP) was read in a spectrophotometer (Biochrom spec Libra S4. Serial no. – 105773, Biochrom Ltd, Cambridge England) at 405 nm and the amount of pNP liberated was calculated by comparing the reading corrected for blanks against a standard curve generated using varying concentrations of pNP. One unit of β–glucosidase activity was defined as the amount of enzyme needed to liberate 1µM of p-Nitrophenol (pNP) per minute under the standard assay conditions.Each experiment was repeated three times and analysed statistically. Standard error of mean (SEM) were calculated using Microsoft excel.
2.2.3. Protein Estimation
- The protein concentration was determined using the Bradford method [41]. Five milliliter of the dye reagent was added to 100 µl of appropriately diluted enzyme sample and was incubated for 5 minutes after which the absorbance was measured at 595 nm in a spectrophotometer (Biochrom spec Libra S4. Serial no. – 105773, Biochrom Ltd, Cambridge England). The amount of protein was determined by extrapolating from a bovine serum albumin (BSA) standard curve, constructed by using a solution containing 1 mg/ml BSA. Bradford protein assay is based on the observation that the absorbance maximum for an acidic solution of Coomasie Brilliant Blue G-250 shifts from 465 nm to 595 nm when binding to proteins occurs. Both hydrophobic and ionic interactions stabilize the anionic form of the dye, causing a visible color change [41].
2.2.4. Enzyme Purification
2.2.4.1. Anion-Exchange Chromatography
- Five milliliter (5 ml) of the dialyzed β-glucosidase sample was loaded onto the DEAE-Sephadex A-50 column equilibrated with phosphate buffer (pH 7). The column was eluted with a linear gradient (0.05M – 0.75 M) NaCl in phosphate buffer at pH 7.0 during washing. The activity and protein concentration of β-glucosidase in each fraction was determined. The active fractions were pooled together, concentrated using polyethylene glycol 1500 (BDH) and dialyzed extensively against phosphate buffer. The homogeneity of the purified β-glucosidase was checked by SDS-PAGE.
2.2.4.2. Determination of Purity and Molecular Mass
- Purity and molecular mass of the partially purified β-glucosidase was determined by SDS-PAGE using an appropriate molecular mass marker. The gel was stained using Sterling rapid silver stain following the manufacturer’s instructions.
2.2.4.3. Effect of pH on β-glucosidase Activity
- The effect of pH on the enzyme activity was studied within the pH 3.0 – 10.0 ranges:10 mM citrate buffer, pH 3.0 – 6.0; 10 mM phosphate buffer, pH 7.0 – 8.0; 10 mM glycine/NaOH buffer, pH 9.0 – 10.0. β-glucosidase activity was then assayed at each pH level. A graph of the β-glucosidase activity versus pH was drawn to determine optimum pH.
2.2.4.4. Effect of Temperature on β-glucosidase Activity
- The effect of temperature on β-glucosidase activity was followed between 20 and 90°C using the optimum pH. A graph of β-glucosidase activity versus temperature was plotted to determine optimum temperature.
3. Results
3.1. Phylogenetic Analysis of β-glucosidases
- A comparison of the phylogenetic analysis of GH1 and GH3 family (Figure 1a) clearly shows GH1 and GH3 sequences forming separate clades. The tree drawn using GH1 and GH3 β-glucosidase sequences from different accessions revealed that the same strain of organism produces both GH1 and GH3 family β-glucosidase, for instance, AFL2G_08626 (belonging to GH1 family) and AFL2G_00957 (belonging to GH3 family) are all from Aspergillus fumigatus.
3.2. Multiple Sequence Alignment of β-glucosidases
- A complete sequence alignment was carried out on the alignment program TCoffee [36] to generate a multiple sequence alignment. The multiple sequence alignment results (Figure 2) revealed that there is good alignment among the β-glucosidase proteins. For instance, AN1804 had 99% identity with 92% coverage to AnBg11 with an E-value of 1e-87. AnBg11 and AN1804 both have signal peptide cleavage at position 19/20 while AN2227.2 and TnBg13B lacked a signal peptide. Multiple sequence alignment also revealed accession TnBg13B to have nine of the core catalytic active sites [43] while AN2227.2 and AN1804.2 had eight each of the core catalytic active sites aligned.
3.3. Small Scale Expression of AN1804 Recombinant β-glucosidase
- A characterization of A. nidulans AN1804 β-glucosidase clones already sub cloned into the expression vector pPICZ [34] was then attempted. Figure 3a shows the changes in crude protein concentration of A. nidulans AN1804 β-glucosidase in Buffered Methanol Complex Medium (BMMY). β-glucosidase protein concentration increased until it reached optima of 0.67 mg/ml after 48 hours and the concentration subsequently declined.
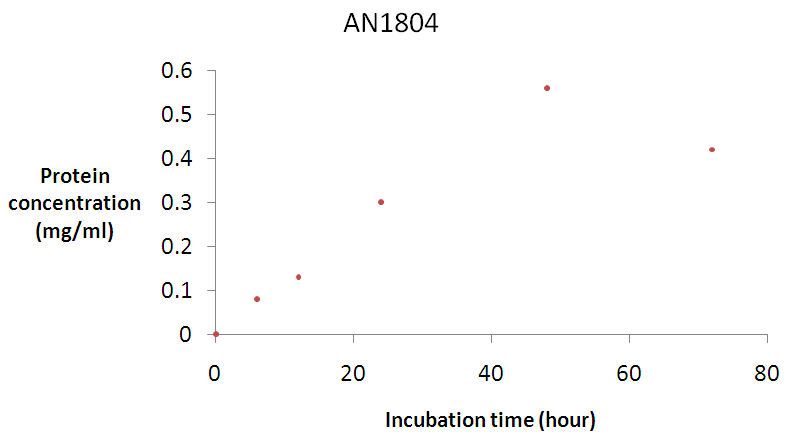 | Figure 3a. Time course for β-glucosidase production by A. nidulans AN1804 Error bars represent the mean ± SEM (n = 3) Error bars represent the mean ± SEM (n = 3) |
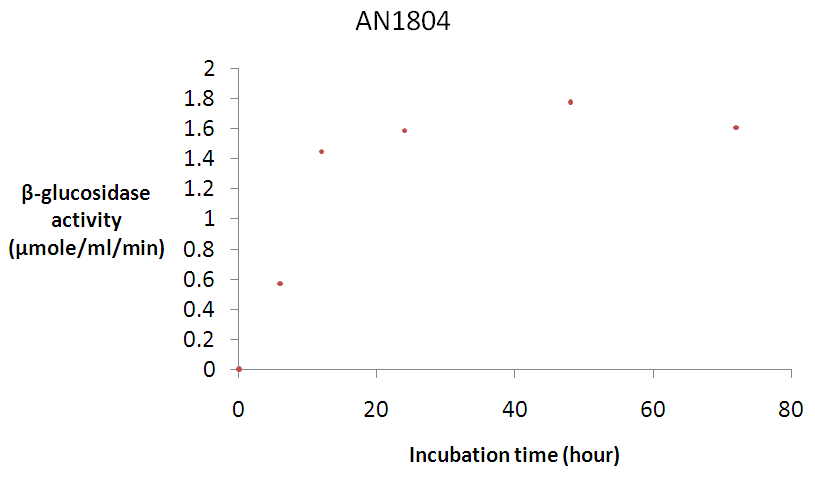 | Figure 3b. Time course for β-glucosidase activity by A. nidulans AN1804 Error bars represent the mean ± SEM (n = 3) Error bars represent the mean ± SEM (n = 3) |
|
3.3.1. Anion-Exchange Chromatography
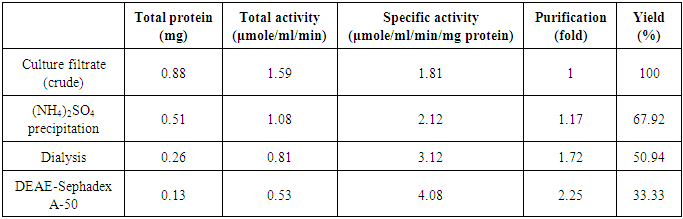
- The anion exchange chromatography elution profile of β-glucosidase is shown in Figure 4. The elution profile resulted in a single broad peak (fractions 10 – 16) which were pulled together and dialysed against the phosphate buffer. The collated fractions produced a final purification of 2.25 fold over the crude extract (Table 1).
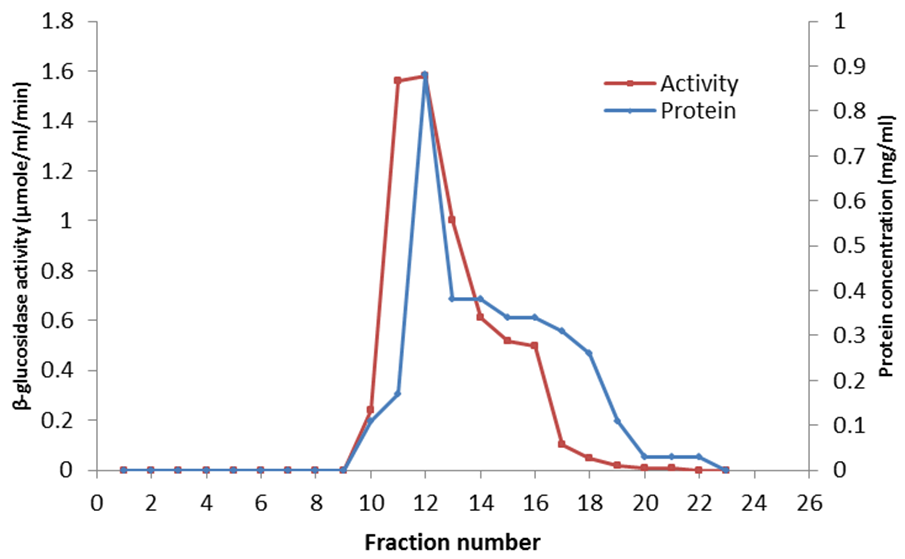 | Figure 4. Elution profile of A. nidulans AN1804 β-glucosidase from DEAE-Sephadex A-50 anion exchange column |
3.3.2. Determination of Purity and Molecular Mass
- The purity and molecular mass of the partially purified β-glucosidase was determined by SDS-PAGE and stained using Sterling rapid silver stain. The purified β-glucosidase enzyme moved homogenously as a single band on the SDS polyacrylamide gel. The β-glucosidase enzyme moved at the same speed corresponding to an estimated molecular weight of 100 kDa as S6-0024 BLUeye prestained protein ladder (Tris-Glycine 4 – 20%) with an estimated concentration mass of 300 ng (data not shown).
3.3.3. Effect of pH on β-glucosidase Activity
- The effect of pH on the β-glucosidase enzyme activity was studied within the pH 3.0 – 10.0 range. Figure 5 shows the pH profile of β-glucosidase AN1804. The enzyme had a strikingly broad pH ranges and was most active at pH 5.5.
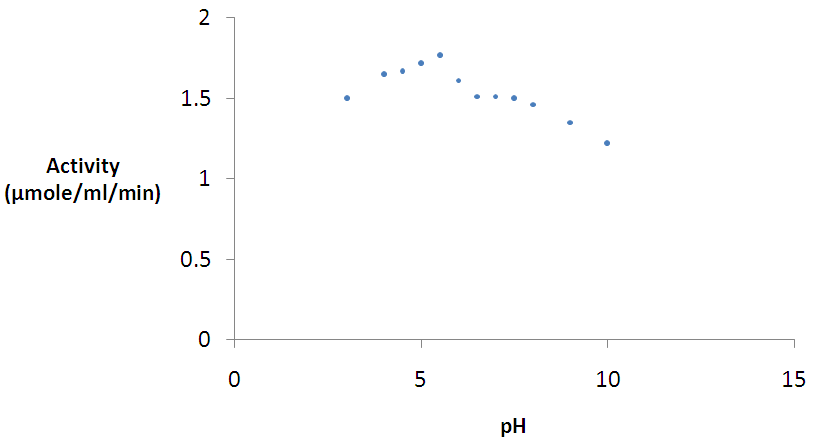 | Figure 5. Effect of pH on β-glucosidase activity Error bars represent the mean ± SEM (n = 3) Error bars represent the mean ± SEM (n = 3) |
3.3.4. Effect of Temperature on β-glucosidase Activity
- Figure 6 shows the temperature profile of β-glucosidase activity. Activity was followed between 20 and 90°C. Using a pH of 5.5 as the optimum pH β-glucosidase activity, the enzyme was optimally active at 50°C (Figure 6). The purified enzyme produced a very broad temperature range of activity and good activity was even observed at 70°C. The activity however dropped sharply at temperatures above 70°C.
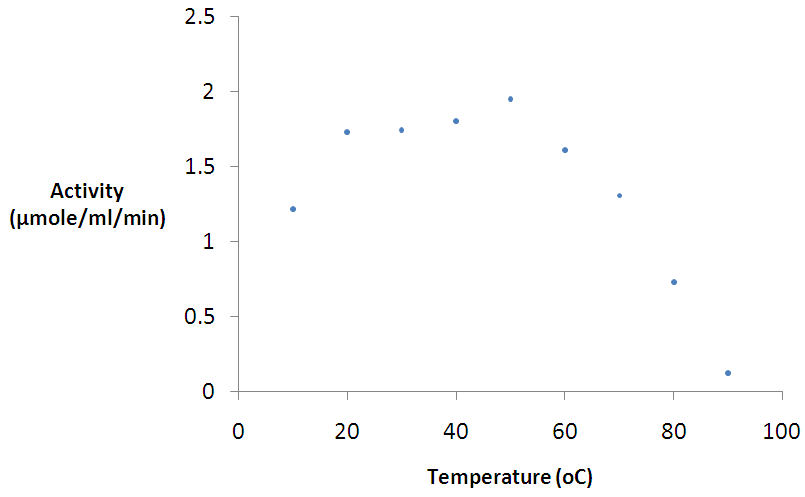 | Figure 6. Effect of temperature on β-glucosidase activity Error bars represent the mean ± SEM (n = 3) Error bars represent the mean ± SEM (n = 3) |
4. Discussion
4.1. Bioinformatics Analysis of β-glucosidase
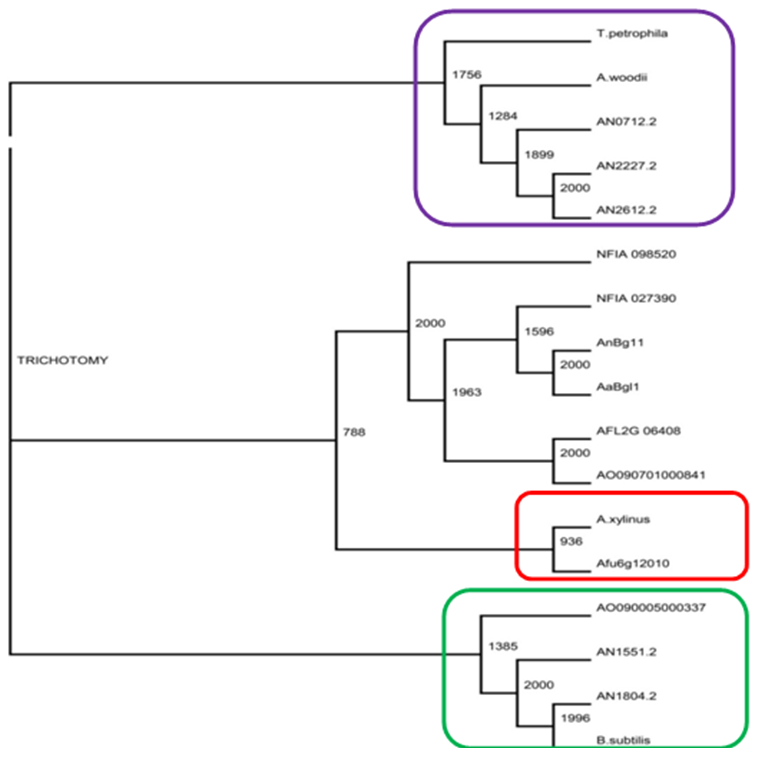
- β-glucosidases act upon β-1→4 bonds linking two glucose molecules and are the key enzyme components present in cellulase that complete the final step during cellulose hydrolysis by converting cellobiose to glucose [24]. They also work in harmony with exoglucanase and endoglucanase to hydrolyse cellulose to fermentable sugars [46, 47]. Phylogenetic tree analysis was carried out to determine evolutionary relationships of AN1804 β-glucosidase proteins with that of bacteria. Figure 1a shows a comparison of the phylogenetic analysis of GH1 and GH3 family sequences clearly forming separate clades. Literature surveys [23, 28] show that two methods are generally used in the classification of β-glucosidases which are based on substrate specificity and nucleotide sequence identity. CAZy classification of GH has placed β-glucosidases in GH family 1 and 3 based on their amino acid sequence structure [22]. Generally, β-glucosidases from bacteria, fungi, plant and mammalian origin belong to GH1 family and usually possess a significant level of galactosidase activity in addition to β-glucosidase activity while β-glucosidases of fungi, bacteria and plants are classified into GH3 family [24, 48]. From a structural point of view, the GH1 family β-glucosidases are said to catalyse substrates following the β-retaining action mechanism that employs glutamic acid as a catalytic nucleophile, while GH3 family utilize an aspartic acid residue in their nucleophile attack of the substrate [43]. The GH1 β-glucosidases consist of proteins with (β/α)8-barrel structures in contrast to the active site of GH3 enzyme which comprises of two domains [49]. Most interestingly, Figure 1b highlights the unexpected association in phylogenetic trees of AN1804 β-glucosidase which is an Aspergillus enzyme with Bacillus subtilis which is a bacterial counterpart, suggesting possible horizontal gene transfer events from bacteria to fungi or convergent evolution. Interestingly, the encoding Aspergillus gene also lacks introns. Crucially the AN1804 enzyme appears to lack a fungal specific extension of amino acids (Figure 2) [43]. Horizontal gene transfer (HGT) in microbes has played an important role in their evolution and in the generation of genes involved in the synthesis of cellulase enzymes which degrade cellulose. The process and mechanisms involved in the exchange of DNA between distant species is unknown [50] but horizontal gene transfer between bacteria and fungi in nature is well documented [51]. Hakkinen et al. [52] showed the horizontal gene transfer of an endoglucanase gene from bacteria (Bacillus subtilis) to fungi (Trichoderma spp.). Another classic example of HGT is documented by Ubhayasekera and Karlsson [53], where Streptomyces genes were shown to have possible homologues in Hypocrea (Trichoderma). In a study on phylogenetic distribution of cellulases in bacteria, Berlemont and Martiny [54] reported that many bacteria (termed opportunistic bacterial strains) produce only β-glucosidases while a small number of potential bacteria cellulose degraders produce both other cellulases and β-glucosidases. Many bacteria therefore act as opportunists, relying on other species with a complete repertoire of cellulose degrading enzymes to provide cellobiose. In contrast Aspergillus species seem to be self-sufficient for cellulose degradation, encoding the full range of enzymes required [22].The present work suggests that AN1804 genes may have acquired specific β-glucosidase genes from bacteria. Evidence for bacterial origin was provided by their lack of introns, absence of some fungal specific amino acid insertions in its protein (Figure 2) sequence, automatic annotation as “periplasmic” and unusual positions in phylogenetic trees showing similarities to bacterial proteins (Figure 1b).
4.2. Small Scale Expression of the Recombinant β-glucosidase
- β-glucosidase AN1804 was purified from Pichia pastoris [34] using Buffered Methanol Complex Medium (BMMY) culture medium. The expression of the enzyme was monitored by screening its activity. Figure 3a shows the time course for crude β-glucosidase production while Figure 3b shows the time course for crude β-glucosidase activity. Although the crude β-glucosidase enzyme produced were found to reach optimum enzyme expression and activity of 0.56 mg/ml and 1.78 μmole/ml/min respectively (Figure 3a and 3b), a lag period was observed before the enzyme activity reached the peak value on the second day following methanol induction. The lag period suggested that there was adsorption of the methanol and as the hydrolysis proceeded, a portion of the adsorbed enzymes was gradually released into the reaction supernatant as suggested by Lee et al. [55]. β-glucosidase was successfully expressed and the enzymes were used for biochemical characterization using pNPG as a substrate.A purification of 1.17 fold was achieved for β-glucosidase after ammonium sulphate precipitation. This result is expected because ammonium sulphate has been reported to be the best protein precipitant without having effect on the biological activity of the enzyme [3]. Ammonium sulphate also has high solubility characteristics and changes of temperature have little or no effect on its solubility [56]. Dialysis produced a purification of 1.72 fold. The increase in purification was due to desalting and the removal of low molecular weight compound.
4.2.1. Anion-Exchange Chromatography
- The extracellular β-glucosidases was purified by ion exchange chromatographuy using DEAE-sephadex A-50. The anion exchange chromatography separation of the enzymes by elution on DEAE-sephadex A-50 columns with linear gradient NaCl (0.05 M – 1.0 M) showed a single broad peak (Figure 4) suggesting the presence of a single active protein in the eluate. The purified enzyme preparation produced the final purification of 2.25 fold with 33.33% recovery of enzyme activity. Kaur et al. [57] reported 4.06 fold purification with 15.89% recovery activity of β-glucosidase isolated from Melanocarpus sp MTCC 3922. The difference in the purification fold and percentage yield of β-glucosidase activity in this work and that reported by Kaur et al. [57] is probably due to the difference in the source of the enzymes. The overall specific activity of β-glucosidase was improved from 1.81 to 4.08. These results indicated that sephadex A-50 column chromatography yielded effectively pure enzyme.Judged by the SDS-PAGE data, the purified β-glucosidases was homogeneous. The molecular weight as estimated by SDS-PAGE was 100 kDa suggesting that the enzyme is monomer. Lima et al. [43] purified a monomeric β-glucosidase from A. niger (AnBgl1) with an apparent molecular weight of 116 kDa. The molecular weight of 100 kDa for A. nidulans AN1804 β-glucosidase correlates with 102 kDA molecular weight reported by Kaur et al. [57] for β-glucosidase from Melanocarpus sp MTCC 3922, and the molecular weight of 91.47 kDa is also reported [58] for recombinant β-glucosidase from Aspergillus fumigatus Z5. Bai et al., [59] also reported a molecular weight of 126 kDa for β-glucosidase isolated from Penicillium simplicissimum-11. Though some β-glucosidases have a simple monomeric structures with around 35 kDa molecular weight, others have dimeric structure with molecular weight of 146 kDa or even trimeric structures with over 450 kDa [60].
4.2.2. Effect of pH and Temperature on β-glucosidase Activity
- The maximum activities for β-glucosidase was observed at pH 5.5. However, the enzyme had a broad acidic pH range and although it was not active in the extreme alkaline pH. The optimal pH of 5.5 for β-glucosidase enzyme activity is near pH 6.0 reported for same enzyme by Bauer et al. [34]. A pH of 4.4 – 5.2 was also reported for β-glucosidases isolated from Penicillium simplicissimumh-11 [59]. The pH optimum reported in this work is also in agreement with those reported for most fungal sources which are near pH 6.0 [34, 57, 61, 62]. Reports have also shown that maximum activity at an acidic pH is a common property of cellulases from fungal sources [63]. However, it is striking that AN1804 shows a very broad pH profile. The effect of temperature on the activity of the purified β-glucosidases on pNPG was also analyzed. The maximum activity was observed at 50°C. A value of 40°C is reported for β-glucosidase from thermophilic fungus Humicola insolens [64] which is lower than that reported for most β-glucosidases; while the value of 50°C reported in this studies fall within the range for most β-glucosidases [57-59]. The optimum temperature was lower than that reported for β-glucosidase from Aspergillus niger KCCM 11239 [65] with an optimum temperature of 70°C. The optimum temperature value of 50°C for β-glucosidase obtained in this work is in agreement with that reported by Bauer et al. [34] for the same enzyme. A possible explanation to why fungal species retain bacterial genes could be because of lifestyle, where the organism kept the gene for adaptation to allow the colonization of new environments and nutritional purposes [51]. However, the most striking characteristics of β-glucosidase AN1804 enzymes are the dramatically broad pH and temperature profiles. So we postulate that AN1804 has been retained by its fungal host as it shows activity across a broad pH and temperature profile. This may allow the new host to access lignocellulose in decaying plant material, by secretion of the enzyme, to access locations at temperatures above its optimum for growth.
5. Conclusions
- A. nidulans is well known for its broad pH growth range, conventionally ascribed to the ability to induce enzymes appropriate to particular external pH values [66]. However, the production of an individual enzyme with both broad pH and temperature profiles would clearly bestow a selective advantage on the host fungus. A key issue addressed in the present study is that the A. nidulans genome may likely holds candidate β-glucosidases that are horizontally transferred from bacteria.
ACKNOWLEDGMENTS
- This work was supported financially by the Petroleum Technology Development Fund (PTDF) of Nigeria. The authors also gratefully acknowledge the Fungal Genetic Stock Centre (FGSC) USA, for supplying the A. nidulans Pichia clone AN1804 that was used in the study.
 Abstract
Abstract Reference
Reference Full-Text PDF
Full-Text PDF Full-text HTML
Full-text HTML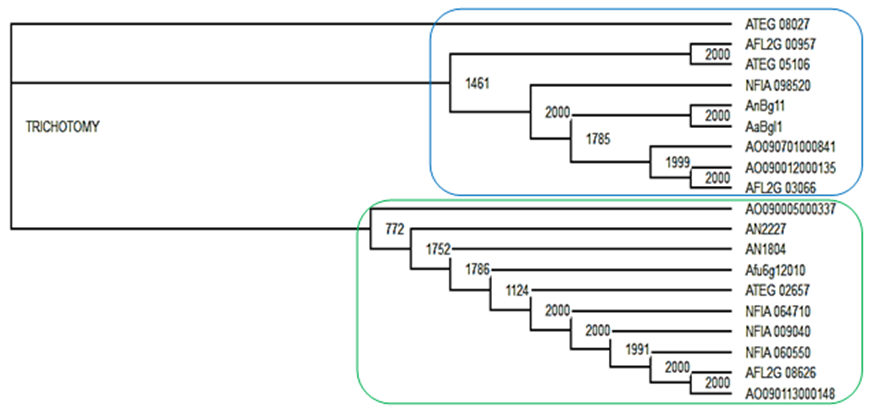

 helix –A and –B (green box), intermediate amino acids from linker 2 and N-terminal domain (black boxes) [43]
helix –A and –B (green box), intermediate amino acids from linker 2 and N-terminal domain (black boxes) [43]